Home > Newsletter Archives
Immune Cells Repurpose a Stem Cell Transcription Factor as a Pathogen Sensor
"Thus, our findings have revealed a previously unknown activation pathway in neutrophils and may provide potential therapeutic strategies to augment the innate immunity of neutrophils for the treatment of infectious diseases," the authors conclude.
Among the most abundant of the immune system's first responders, neutrophil granulocytes comprise up to 75 percent of the body's white blood cells. During the acute phase of inflammation, neutrophils migrate to the site of injury or infection through the circulatory system and interstitial tissues to identify and ingest pathogens. They also release proinflammatory cytokines and chemokines which recruit other immune cells to the inflammatory site to eliminate foreign bacteria.
In order to identify pathogens, neutrophils express toll-like receptors (TLRs) for the recognition of bacterial elements. Recently, a group of researchers from the Chinese Academy of Sciences investigating TLRs determined that neutrophils actually use a transcription factor called Sox2, already known for its role in maintaining the self-renewal of ectodermal and endodermal stem cells, as one such TLR.
Observing that the morphology of Sox2 present in stem cells resembled that of neutrophils, the researchers determined through immunohistochemical staining that mouse neutrophils, like stem cells, also stained with the antibody for Sox2. But they did not find the Sox2 transcription factor in other cells, including T cells, B cells, natural killer cells, dendritic cells, macrophages, or other granulocytes. "These data indicated that among mature hematopoietic cells, only neutrophils expressed Sox2," the authors write.
The function of Sox2 in immune response
In order to explore the physiological role of Sox2 in the immune system, the researchers bred mice with the conditional deletion of Sox2 in neutrophils; littermates with normal expression of Sox2 served as controls. Both sets of mice were infected with Listeria monocytogenes to determine if Sox2 played an in vivo role in clearing bacteria.
The results were stark, as the Sox2-deficient mice died at a higher rate than their littermate counterparts, and exhibited a bacterial load in the spleen and liver 10,000-fold higher than the control group. This, along with corroborating data, suggest a critical role for Sox2 in neutrophil-mediated bacterial clearance.
Neutrophils require Sox2 to identify bacterial DNA
To test the hypothesis that neutrophils use Sox2 transcription factor as a sensor to identify bacterial DNA, the researchers isolated neutrophils from both sets of mice and transfected the samples in vitro with bacterial genomic DNA. The proinflammatory responses of Sox2-deficient neutrophils were 10 times lower than the unmodified neutrophils.
They also explored Sox2 binding sequences in the Listeria genome, determining that 90 percent of sequences that matched Sox2 would bind to it. Furthermore, an analysis of the genomes for other bacteria, including Bartonella, Staphylococcus, and Salmonella found sequences matching Sox2, and which, when transfected into normal neutrophils, produced a proinflammatory response. Unmatched DNA fragments used as controls were unable to bind with Sox2. The authors conclude, "Overall, these results indicated that Sox2 sensed bacterial DNA in neutrophils in a sequence-specific manner."
Furthermore, the researchers determined that, though Sox2 functions as a transcription factor in stem cells, its role in neutrophils is exclusively for the identification of foreign DNA. Though a variety of DNA sensors have been identified, the researchers believe that Sox2 plays a pivotal role in the recognition of pathogens and in distinguishing genetic "self" and "non-self" in order to prevent damage to host cells during immune response. "Thus, our findings have revealed a previously unknown activation pathway in neutrophils and may provide potential therapeutic strategies to augment the innate immunity of neutrophils for the treatment of infectious diseases," the authors conclude.
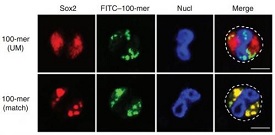
Illustration: Confocal microscopy of Sox2 in wild-type neutrophils transfected for 6 h with fluorescein isothiocyanate (FITC)-conjugated 100-nucleotide matched or unmatched DNA. Dashed outline (far right) indicates neutrophil profile. Scale bar, 5 µm. Nature Immunology (2015) doi:10.1038/ni.3117.
Read more
MedicalXpress (03/05/15)
Abstract (Nature Immunology; 16, 366-375 (03/02/15))
In order to identify pathogens, neutrophils express toll-like receptors (TLRs) for the recognition of bacterial elements. Recently, a group of researchers from the Chinese Academy of Sciences investigating TLRs determined that neutrophils actually use a transcription factor called Sox2, already known for its role in maintaining the self-renewal of ectodermal and endodermal stem cells, as one such TLR.
Observing that the morphology of Sox2 present in stem cells resembled that of neutrophils, the researchers determined through immunohistochemical staining that mouse neutrophils, like stem cells, also stained with the antibody for Sox2. But they did not find the Sox2 transcription factor in other cells, including T cells, B cells, natural killer cells, dendritic cells, macrophages, or other granulocytes. "These data indicated that among mature hematopoietic cells, only neutrophils expressed Sox2," the authors write.
The function of Sox2 in immune response
In order to explore the physiological role of Sox2 in the immune system, the researchers bred mice with the conditional deletion of Sox2 in neutrophils; littermates with normal expression of Sox2 served as controls. Both sets of mice were infected with Listeria monocytogenes to determine if Sox2 played an in vivo role in clearing bacteria.
The results were stark, as the Sox2-deficient mice died at a higher rate than their littermate counterparts, and exhibited a bacterial load in the spleen and liver 10,000-fold higher than the control group. This, along with corroborating data, suggest a critical role for Sox2 in neutrophil-mediated bacterial clearance.
Neutrophils require Sox2 to identify bacterial DNA
To test the hypothesis that neutrophils use Sox2 transcription factor as a sensor to identify bacterial DNA, the researchers isolated neutrophils from both sets of mice and transfected the samples in vitro with bacterial genomic DNA. The proinflammatory responses of Sox2-deficient neutrophils were 10 times lower than the unmodified neutrophils.
They also explored Sox2 binding sequences in the Listeria genome, determining that 90 percent of sequences that matched Sox2 would bind to it. Furthermore, an analysis of the genomes for other bacteria, including Bartonella, Staphylococcus, and Salmonella found sequences matching Sox2, and which, when transfected into normal neutrophils, produced a proinflammatory response. Unmatched DNA fragments used as controls were unable to bind with Sox2. The authors conclude, "Overall, these results indicated that Sox2 sensed bacterial DNA in neutrophils in a sequence-specific manner."
Furthermore, the researchers determined that, though Sox2 functions as a transcription factor in stem cells, its role in neutrophils is exclusively for the identification of foreign DNA. Though a variety of DNA sensors have been identified, the researchers believe that Sox2 plays a pivotal role in the recognition of pathogens and in distinguishing genetic "self" and "non-self" in order to prevent damage to host cells during immune response. "Thus, our findings have revealed a previously unknown activation pathway in neutrophils and may provide potential therapeutic strategies to augment the innate immunity of neutrophils for the treatment of infectious diseases," the authors conclude.
Illustration: Confocal microscopy of Sox2 in wild-type neutrophils transfected for 6 h with fluorescein isothiocyanate (FITC)-conjugated 100-nucleotide matched or unmatched DNA. Dashed outline (far right) indicates neutrophil profile. Scale bar, 5 µm. Nature Immunology (2015) doi:10.1038/ni.3117.
Read more
MedicalXpress (03/05/15)
Abstract (Nature Immunology; 16, 366-375 (03/02/15))
Newsletter Archives
U.S. News
World News
Regenerative Medicine Journal
Point Of View
World News
Researchers Discover How to Overcome a Treatment Resistance Mechanism in One of the Most Aggressive Types of Breast Cancer Flatworm-Inspired Medical Adhesives Stop Blood Loss Tumors Form Temporary Structures to Avoid Immunotherapy Treatments Epigenetic Treatment in Mice Improves Spinal Cord Regeneration After Injury, Study Shows The Physics of the Premature Lung: Why Mechanical Ventilation Can Harm Preterm Lungs T Cells Use Force to Destroy Cancer Cells Immunotherapy Reduces Lung and Liver Fibrosis in Mice Nerve Healing: Neighboring Cells Become Police Force -- And Could Make Tumors Benign An Unstable, Flake-Like Network in the Making Pioneering Research Using Bacteria Brings Scientists a Step Closer to Creating Artificial Cells with Lifelike Functionality From Wound Healing to Regeneration One in Four Children Who Have Suffered a Minor Head Injury Is Liable to Suffer from Chronic Post-Concussion Syndrome Stretchable Extracellular Matrix Membrane For In Vitro Barrier Models Bacterial Enzyme Makes New Type of Biodegradable Polymer Study Identifies New Protection Mechanism in Breast Cancer Engineers Develop a Magnetic Tentacle Robot to Pass into the Narrow Tubes of the Lung Lung Tissue from the Lab Immune Cells Linked to Neurological Disease Prognosis and Survival Cell Fusion Awakens Regenerative Potential of Human Retina Nano Carriers Fit for Purpose The Cellular Cleaning Program Autophagy Helps in Wound Healing Lab-Grown Pig Heart Tissue Could Help Replace Live Animals in Heart Disease Research Migrating Through Small Spaces Makes Cancer Cells More Aggressive A Speed Limit Could Be a Breakthrough for Stem Cell Therapy New Modelling Framework Developed to Improve Infectious Disease Control Protect Against Aging and Age-Related Disease with the MondoA Protein A Cellular Stress State Predicts a Poor Chemotherapy Response in Ovarian Cancer Patients Stem Cell Signaling: Molecular Morse Code in Stem Cells Encrypting Differentiation Information RNA Molecules Control Repair of Human DNA in Cancer Cells Make No Bones About It: How Our Bodies Shift Between Making Bones and Breaking Them Down Connecting Science to Medicine: Tendon-Like Tissue Created from Human Stem Cells Building Artificial Nerve Cells Sonic Advance: How Sound Waves Could Help Regrow Bones 3D Model of Living Brain Cancer Points to Possible Future for Drug Screening Metal Mix and Match: An Unexpected Discovery Could Improve the Crystallinity of Coordination Nanosheets Self-Healing Materials for Robotics Made from Jelly and Salt Nebulin No Longer Nebulous! Scientists Obtain First High-Resolution 3D Image of Muscle Protein Treatment for Parkinsons Could Now Get Even Better MRI Sheds Light on COVID Vaccine-Associated Heart Muscle Injury Researchers Find Trove of Proteins that May Influence Cystic Fibrosis Highly Responsive Immune Cells Seem to Be Beneficial for the Brain Potential to Identify Risk of Alzheimers in Middle Age with Simple Eye Test Colonic Gene Mapping Gives Insights into Intestinal Diseases New Implant Offers Promise for the Paralyzed A New Multipurpose On-Off Switch for Inhibiting Bacterial Growth Study in Mice Shows Potential for Gene-Editing to Tackle Mitochondrial Disorders Suppressing the Spread of Tumors Important Step Towards Fasting-Based Therapies Sweet Pressure: Scientists Discover Link Between High Blood Pressure and Diabetes Complex Three-Dimensional Kidney Tissue Generated in the Lab from Scratch Protein Machinery of Respiration Becomes Visible 'Boot Camp' Enzyme Prevents Autoimmune Conditions Lucky Genes Can Help Protect People with Obesity from Some Disease Patient-Derived Heart Cells Mimic Disease In Vitro Artificial Pancreas Proves Life-Changing for Very Young Children with Type 1 Diabetes and Their Families Protective Gene Variant Against COVID-19 Identified More Insight into How Vision Works Parkinsons Protein Blueprint Could Help Fast-Track New Treatments Developing Bioactive Coatings for Better Orthopaedic Implants Is Energy the Key to Alzheimers Disease? Innovative X-Ray Imaging Shows COVID-19 Can Cause Vascular Damage to the Heart Radiation Could Damage Biological Tissue Also Via a Previously Unnoticed Mechanism Link Between Long-Term Exposure to Air Pollution and Fatty Liver Disease Shown Researchers Crack the Synthetic Code of Rare Molecules Sought After in Drug Development Printing Technique Creates Effective Skin Equivalent, Heals Wounds Long-Range Four-Stranded DNA Structures Found to Play a Role in Rare Aging Disease Genes Associated with Hearing Loss Visualized in New Study Synthetic Tissue Can Repair Hearts, Muscles, and Vocal Cords Gold-Based Cancer Therapy Could Face Competition from Other Substances New Link Between Diet, Intestinal Stem Cells, and Disease Discovered Using Mussel Adhesive Proteins for Cardiac Tissue Regeneration Extreme Heat Events Jeopardize Cardiovascular Health, Experts Warn Using T Cells to Target Malignant Brain Tumors Microbiome Discovery May Open New Doors to Development of Treatments for Gastrointestinal Diseases Lack of Important Molecule in Red Blood Cells Causes Vascular Damage in Type 2 Diabetes Microbiome Discovery Could Help Save Kids Hearing Bone Marrow Backup Needed to Tackle Respiratory Infections Giant Leap Taken in Fighting Antibiotic Resistance New Test to Diagnose Aggressive Childhood Brain Tumors Cause of Alzheimers Progression in the Brain How COVID-19 Alters the Immune System Computer Scientists Developed Method for Identifying Disease Biomarkers with High Accuracy Flexible Device Could Treat Hearing Loss Without Batteries Promising New Antimalarial Compound Discovered Lab-Grown Mini Brains Hint at Potential Treatment Options for Motor Neuron Disease and Frontotemporal Dementia A Mathematical Model to Help Optimize Vaccine Development Our Brains Have a 'Fingerprint' Too New Technique Helps Researchers Understand How Acid Damages Teeth Gel Fights Drug-Resistant Bacteria and Induces Bodys Natural Immune Defense Filling The Gaps: Connecting Genes to Diseases Through Proteins Mutated Cells Drive Out Early Tumors from the Esophagus Colorblind Fish Show Experts How Vision Evolved Novel Fluorescence Imaging Assay Gives New Insight to Develop Biomimetic Nanoparticles for Targeted Cancer Therapy Super-Enhancers: The Villain Fueling Certain Cancers Link Between Crohns Disease and Fatty Tissue in the Gut Revealed The Immune System's Double Agents Directly Into the Brain: A 3D Multifunctional and Flexible Neural Interface Antidepressants Inhibit Cancer Growth in Mice Gut Bacteria Influence Brain Development Discovery Highlights the Complex Lifestyles of Frontline Immune Cells When Organoids Meet Coronaviruses How Resistant Germs Transport Toxins at Molecular Level Faster Research Through Automation: New Way to Count Micronuclei in Cells Likely Cause of Alzheimers Disease Identified New Findings on B Cells May Improve Vaccine Design Researchers Improve Optical Tissue Clearing Method to Diagnose Cancer Newly Developed Software Unveils Relationships Between RNA Modifications and Cancers New Research Shows a Link Between Cell Identities and Childhood Cancer Type Neuroblastoma Over 200 Health Journals Call on World Leaders to Address 'Catastrophic Harm to Health' from Climate Change New Resource to Advance Genomics-Driven Precision Medicine Predicting Possible Alzheimers with Nearly 100 Percent Accuracy Fighting Brain Cancer at Its Root How Can I Avoid Heart Disease or Stroke? AI Helps to Spot Single Diseased Cells Increased Survival with Eye Melanoma in Clinical Trial Mathematical Model Predicts Best Way to Build Muscle New Imaging Tool Visualizes Cell Functions in a Microphysiological System Does Alzheimers Disease Start Inside Nerve Cells? Synthetic Tissue Model with Blood Vessels New Insights into How the 'First Brain' Works in the Gut A Defective Potassium Channel Disrupts the Brains Navigation System Getting Oxygenated Blood to Exercising Muscles Graphene Binds Drugs Which Kill Bacteria on Medical Implants New CRISPR/Cas9 Technique Corrects Cystic Fibrosis in Cultured Human Stem Cells Scientists Reverse a Key Hallmark of Motor Neuron Disease in the Laboratory New Drug Molecules Could Prevent Skin Aging Caused by Sun Exposure Newly Developed, Bioinspired Cell Delivery Vehicles New Breakthrough to Help Immune Systems in the Fight Against Cancer A Novel Method for the Rapid Repair of Peripheral Nerve Injuries New Understanding of Cell Stability with Potential to Improve Immune Cell Therapies 'Missing Self' Contributes to Organ Rejection After Transplantation Scientists Reverse Age-Related Memory Loss in Mice Why Identical Mutations Cause Different Types of Cancer Glaucoma Test 'Best Yet' Metal-Based Molecules Show Promise Against the Build-Up of Alzheimer's Peptides Potential Marker for Success of Immunotherapy in the Treatment of Lung Cancer Bone Marrow-Derived Fibrin Clot as Better Source for Meniscal Repair An Antioxidative Stress Regulator Protects Muscle Tissue in Space, Mouse Study Shows Shortcut for Dendritic Cells New Epigenetic Regulatory Mechanisms Involved in Multiple Myeloma Growth Cells from the Center of Tumors Most Likely to Spread Around the Body Simple Surgery Prevents Strokes in Heart Patients New Immunotherapy 'Highly Effective' Against Hepatitis B Research Results Challenge a Decades-Old Mechanism of How We Hear Sounds Managing Children's Weight, Blood Pressure and Cholesterol Protects Brain Function Mid-Life Distinct Cell-To-Cell Communication Processes Controlled Differently Why Is COVID-19 So Hard to Treat? Growing Evidence Points to Unique Infectious Profile Engineering Diseased Human Skin In Vitro Thin and Brittle Bones Strongly Linked to Women's Heart Disease Risk Feeling Younger Buffers Older Adults from Stress, Protects Against Health Decline Insights from Color-Blind Octopus Help Fight Human Sight Loss Scientists Invent Catheter System to Deliver Electricity-Activated Glue Path A Physics Perspective on Wound Healing A New Way of Rapidly Counting and Identifying Viruses Muscle Gene Linked to Type 2 Diabetes Force Transmission Between Cells Orchestrates Collective Cellular Motion Newly Discovered Immune Cell Function Vital to Healing Inspired by Nature, the Research to Develop a New Load-Bearing Material Defects in a Specific Cell Type May Cause Ulcerative Colitis Men's Loneliness Linked to an Increased Risk of Cancer Biologists Investigate Effects of Bisphenols on Nerve Cells 165 New Cancer Genes Identified with the Help of Machine Learning A Tummy Invader: This Bacterial Molecule May Be Key to Fighting Stomach Cancer Possible Trigger for Crohn's Disease Identified Vitamin A for Nerve Cells Scientists Use Nanotechnology to Detect Bone-Healing Stem Cells New Genetic Clues Point to New Treatments for 'Silent' Stroke AI Used in Battle Against Asbestos-Linked Cancer Deluge of DNA Changes Drives Progression of Fatal Melanomas Bacteria May Aid Anti-Cancer Immune Response The Astonishing Self-Organization Skills of the Brain Treatment for Type-2 Diabetic Heart Disease New Compound Targets Enzyme Linked to Autoimmune Disorders, Severe COVID-19 Insulin Rises Before Cells Develop Resistance, New Diabetes Research Implies Predicting Success in Therapy with Individualized Cancer Models Immune Cells in Cerebrospinal Fluid Predict Response to Immunotherapy New Method Facilitates Development of Antibody-Based Drugs Tackling Tumors with Two Types of Virus Searching for Novel Targets for New Antibiotics Could Our Immune System Be Why COVID-19 Is So Deadly? Acute Breakdown of the Glial Network in Epilepsy Biobattery-Powered Microneedle Patch Can Deliver Drugs and Procure Testing Samples Fat Cells May Influence How the Body Reacts to Heart Failure Bioengineered Hybrid Muscle Fiber for Regenerative Medicine Advanced Imaging Technology Captures Translation of the Maternal Genome Improved Vectors for Ocular Gene Therapy Making Sense of the Mass Data Generated from Firing Neurons Mitochondrial Function Can Play Significant Part in Serious Disease Scientists Identify More Than 140,000 Virus Species in the Human Gut Potential New Treatment for Fatal Childhood Brain Cancer Scientists Identify How Harmless Gut Bacteria 'Turn Bad' New Research Identifies Biological Causes of Muscle Weakness in Later Life Monash University Novel Protein Could Reverse Severe Muscle Wasting in Disease, Aging and Trauma New Insight into Protein Structures that Could Treat Huntington's Disease Researchers Have Broken the Code for Cell Communication Inherited Immune Condition Reversed by Random DNA Change Scientists Discover a New Pathway Essential for Blood Formation Can Large Fluid-Filled Spaces in the Brain Help Identify Those at Risk of Dementia? New Control Mechanism in Innate Immune System New Technique to Fast-Track Pain Research Air Pollution Linked to Higher Risk of Sight Loss from AMD Breakthrough Design Vastly Improves Mechanical Heart Valve Wirelessly Rechargeable Soft Brain Implant Controls Brain Cells Gut Microbiota Reveals Whether Drug Therapies Work in Inflammatory Bowel Diseases Scientists Use a Novel Ink to 3D Print 'Bone' with Living Cells Designer Cytokine Makes Paralyzed Mice Walk Again New Clues to Prostate Cancer SARS-Cov-2 Antibody Test Helps Select Donor Blood Samples for Therapeutic Use Uncovering Basic Mechanisms of Intestinal Stem Cell Self-Renewal and Differentiation A Brain Mechanism Underlying 'Vision' in the Blind Is Revealed Anticoagulants Reduce the Number of Brain Metastases in Mice Discovery of Aging Mechanism for Hematopoietic Stem Cells New Research May Explain Severe Virus Attacks on the Lungs Global Trial Reveals Life Saving Drug for Acute Myeloid Leukemia Stopping RAS Inhibitors Tied to Worse Outcomes in Patients with Chronic Kidney Disease A New Means of Neuronal Communication Discovered in the Human Brain Nanotechnology: Nanoparticles as Weapons Against Cancer Scientists Build Whole Functioning Thymus from Human Cells Muscle Cell Secrets tRNA Fragments Are Involved in Post-Stroke Immune Reactions Researchers Suggest Stool Transplants Can Battle Serious Infections Gut Microbiota Plays a Role in Brain Function and Mood Regulation Useful 'Fake' Peptides New DNA Modification 'Signature' Discovered in Zebrafish Findings About Cilia on Cells of the Vessel Wall May Be Relevant for Diabetes Treatment Small Molecules Control Bacterial Resistance to Antibiotics Researchers Develop New Biomaterial that Helps Bones Heal Faster Gut Microbes: A Key to Normal Sleep Scientists Develop New Gene Therapy for Eye Disease Simulations Open a New Way to Reverse Cell Aging Cutting Edge Technology to Bioprint Mini-Kidneys Potential New Target to Combat Inflammatory Diseases Biofriendly Protocells Pump Up Blood Vessels Breakthrough in Childhood Brain Cancer Will Save Lives COVID-19 Survival Among Elderly Patients Could Be Improved by Arthritis Drug, Study Finds Predicting the Risk of Severe Side Effects of Cancer Treatment Printable Ink Guides Cell Growth, Offers Nerve Injury Hope New Technique May Revolutionize Accuracy and Detection of Biomechanical Alterations of Cells Repetitive Elements Trigger RIG-I-Like Receptors to Enhance Hematopoietic Stem Cell Formation DNA Repair Supports Brain Cognitive Development Shining a Light on the Role of the Genome's 'Dark Matter' in Cancer Development 90% Accuracy in Detecting Melanoma: New Non-Invasive Method Mystery Molecule in Bacteria Is Revealed to Be a Guard Machine Learning Predicts Anti-Cancer Drug Efficacy Healthcare App Reduces Symptoms of COPD Compared to Regular Treatment A Malformation Illustrates the Incredible Plasticity of the Brain How the Immune System Remembers Viruses A Groundbreaking Genetic Screening Tool for Human Organoids New Estimates of Breast Cancer Risks Associated with HRT The Protein Dress of a Neuron Tiny Golden Bullets Could Help Tackle Asbestos-Related Cancers Biomarkers Could Be Used in a Quick, Inexpensive COVID-19 Blood Screening Tool New Tactic to Stop the Growth of a Deadly Brain Cancer New Therapeutic Approach Against Leukemia Study Explains the Process that Exacerbates MS How Cancer Cells Escape Crowded Tumors Bone Loss: Perforated Bone Tissue from Too Little Sugar Breakthrough Discovery in Gene Causing Severe Nerve Conditions Hand-Held Device Reads Levels of Cancer Biomarker New Class of Highly Effective Inhibitors Protects Against Neurodegeneration New Key Player in Long-Term Memory Coupling Antibiotics with Stem Cells to Fight Off Bone Infections Millimeter-Precision Drug Delivery to the Brain A Small Switch with a Big Impact New Tool Shows Main Highways of Disease Development Silk Fibers Improve Bioink for 3D-Printed Artificial Tissues and Organs Complex Interplay Among Cells Guides Them to Where They Need to Go World First Study Links Obesity with Reduced Brain Plasticity Carriers of Two Genetic Mutations at Greater Risk for Illness and Death from COVID-19 Breakthrough for Tomorrow's Dentistry Is Rheumatoid Arthritis Two Different Diseases? Parkinson's Disease Is Not One, But Two Diseases A Link Between Sensory Neurons Activation and the Immune System Improving the Efficacy of Cellular Therapies A Safer UV Light Effectively Kills Virus Causing COVID-19, Study Shows Painless Paper Patch Test for Glucose Levels Uses Microneedles Fast and Efficient Method to Produce Red Blood Cells Developed New Immunotherapy to Beat Cancer Stem Cell Research Delivers New Points of Attack Against Parkinson's Disease The Genetics of Blood: A Global Perspective Unconventional T Cells in Severe COVID-19 Patients Could Predict Disease Outcome Novel Insights of How Prostate Cancer Causes Secondary Tumors Elderly People Protected Against Respiratory Infections by BCG Vaccine What Happens When Babies with Heart Defects Become Adults? Giant Nanomachine Aids the Immune System Preventing Infection, Facilitating Healing: New Biomaterials from Spider Silk Synthetic Compound Could Serve as Prototype for Novel Class of Drugs to Treat Neurological Damage Antibody Blockade Effective in Treatment of Severe COVID-19 Reproducing the Pathophysiology of Polycystic Kidney Disease from Human iPS Cells Mechanisms Identified to Restore Myelin Sheaths After Injury or in Multiple Sclerosis Fat Crystals Trigger Chronic Inflammation New Insights into How Skin Can Regenerate After Severe Burns How Protein Protects Against Fatty Liver Immune Response to Sars-Cov-2 Following Organ Transplantation Simultaneous Stimulation Helped a Spinal Cord Injury Patient Regain the Ability to Walk Tumor Gene Test Could Help to Predict Ovarian Cancer Prognosis Newly Identified Gut Cells Nurture Lymph Capillaries Becoming a Nerve Cell: Timing Is of the Essence Chemist's New Process Fast-Tracks Drug Treatments for Viral Infections and Cancer New Strategy Against Osteoporosis Move Over Akita: Introducing 'Kuma Mutant' Mice for Islet Transplantation Research Hydrogel Paves Way for Biomedical Breakthrough Scientists Develop New Way to Deliver More Drugs Through the Skin The Enemy Within: Safeguarding Against the Spread of Intracellular Bacteria Tinkering with Roundworm Proteins Offers Hope for Anti-Aging Drugs New Drug Targets for Lethal Brain Cancer Discovered New Drug Targets for Lethal Brain Cancer Discovered Compounds Show Promise in Search for Tuberculosis Antibiotics Peering into the Secrets of Phages to See How They Kill Bacterial Superbugs Unparalleled Inventory of the Human Gut Ecosystem Different from a Computer: Why the Brain Never Processes the Same Input in the Same Way Simple Test Helps to Predict and Prevent Falls UK's Vaccine Against SARS-Cov-2 Is Safe and Induces an Immune Reaction, According to Preliminary Results Chinese Phase 2 Trial Finds COVID-19 Vaccine Is Safe and Induces an Immune Response Multidisciplinary Approach More Effective for Gut Disorders A Novel Therapeutic Target for Recovery After Stroke Age-Related Impairments Reversed in Animal Model Neurobiology: How Much Oxygen Does the Brain Need? Blood Tests Can Predict the Risk of Liver Cirrhosis Novel Pathology Could Improve Diagnosis and Treatment of Huntington's and Other Diseases Novel Protein Drives Cancer Progression Implants: Can Special Coatings Reduce Complications After Implant Surgery? Promising Therapy for Cardiac Regeneration Clostridium Difficile: Fecal Microbial Transplantation More Effective and Less Costly than Antibiotics Breakthrough Discovery to Transform Prostate Cancer Treatment Entry Point for Curbing the Evolution of Antibiotic Resistance Discovered New Light Shone on Inflammatory Cell Death Regulator Fantastic Muscle Proteins and Where to Find Them Renewed Hope for Treatment of Pain and Depression Physical Activity Prevents Almost 4 Million Early Deaths Worldwide Each Year Shining Light on a Malignant Lung Cancer Muscles Support a Strong Immune System Brothers in Arms: The Brain and Its Blood Vessels A Newly Discovered Disease May Lead to Better Treatment of Cystic Fibrosis Restoring Vision by Gene Therapy New Research into Stem Cell Mutations Could Improve Regenerative Medicine Cancer Cells Cause Inflammation to Protect Themselves from Viruses New Method to Map Cholesterol Metabolism in Brain How Toxic Protein Spreads in Alzheimer's Disease Molecular Pair Offers Potential for Parkinson's Treatment The Fight Goes On: Clinical Trial Shows Promising New Treatment for Rare Blood Cancer Optimal Time to Treat Huntington's Disease Identified First Human Trial of COVID-19 Vaccine Finds It Is Safe and Induces Rapid Immune Response Implantable Biosensor Operates Without Batteries Artificial Pieces of Brain Use Light to Communicate with Real Neurons Designing Vaccines from Artificial Proteins Releasing Molecular 'Brake' Kick-Starts Immune Cell Function New Bone-Graft Biomaterial Gives Patients a Nicer Smile and Less Pain Further Evidence Does Not Support Hydroxychloroquine for Patients with COVID-19 One-Two Punch May Help Fight Against Salmonella A Disease Trigger for Pancreatitis Has Been Identified World-First Saliva Test Detects Hidden Throat Cancer Single-Cell RNA Seq Developed to Accurately Quantify Cell-Specific Drug Effects in Pancreatic Islets New AI Diagnostic Can Predict COVID-19 Without Testing More Selective Elimination of Leukemia Stem Cells and Blood Stem Cells Accumulation of Gene Mutations in Chronic Graft-Versus-Host Disease New Understanding of Asthma Medicines Could Improve Future Treatment COVID-19: The Downside of Social Distancing A Cellular Mechanism Protecting Against Cancer How Hearing Loss in Old Age Affects the Brain A New Therapeutic Target Turns the Immune System Against Lymphoma A New Biosensor for the COVID-19 Virus AI to Make Dentists' Work Easier Milestone for the Early Detection of Sepsis Scientists Identify Cause of Leakiness in Eye Diseases CRISPR Helps Identify Potential Alzheimer's-Related Protein Bile Metabolite of Gut Microbes Boosts Immune Cells Neural Circuits Mapped: Now We Understand Vision Better Keratin Scaffolds Could Advance Regenerative Medicine and Tissue Engineering for Humans Switching On a Key Cancer Gene Could Provide First Curative Treatment for Heart Disease Experience Matters for Immune Cells Repairing Stroke-Damaged Rat Brains Diagnostics, Meet CRISPR Ear's Inner Secrets Revealed with New Technology What Type of Cells Does the Novel Coronavirus Attack? Cell Muscle Movements Visualized for First Time Tissue Dynamics Provide Clues to Human Disease Trial Drug Can Significantly Block Early Stages of COVID-19 in Engineered Human Tissues Air Pollution Linked to Dementia and Cardiovascular Disease Completely New Antibiotic Resistance Gene Has Spread Unnoticed to Several Pathogens Cellular Train Track Deformities Shed Light on Neurological Disease Keeping Lower Back Pain at Bay: Better Exercises Too Much Sugar Leads to Early Death, But Not Due to Obesity Graphite Nanoplatelets on Medical Devices Kill Bacteria and Prevent Infections Stem Cells and Nerves Interact in Tissue Regeneration and Cancer Progression Stroke: When the System Fails for the Second Time Nanostructured Rubber-Like Material with Optimal Properties Could Replace Human Tissue Parkinson's Disease Linked to Gene Targeted by Blue-Green Algae Toxin Gold Nanoparticles Uncover Amyloid Fibrils How Intermittent Fasting Changes Liver Enzymes and Helps Prevent Disease AI Taps Human Wisdom for Faster, Better Cancer Diagnosis Seismic Imaging Technology Could Deliver Finely Detailed Images of the Human Brain Bacteria Killed by New Light-Activated Coating New Sleep Method Strengthens Brain's Ability to Retain Memories Biomaterial Discovery Enables 3D Printing of Tissue-Like Vascular Structures New Study Unveils the Mechanism of DNA High-Order Structure Formation The First Cell Atlas for the Human Thymus Sugar Gets the Red Light from Consumers in New Study Immunology: Tonsils as a Testbed DNA Discovery Can Lead to New Types of Cancer Drugs Two Sides of a Coin: Our Own Immune Cells Damage the Integrity of the Blood-Brain Barrier GPS for Chromosomes: Reorganization of the Genome During Development 'Resetting' Immune Cells Improves Traumatic Brain Injury Recovery in Preclinical Trials Leukemia Drugs Hold Promise for Treatment-Resistant Lung Cancer Origins of Immune System Mapped, Opening Doors for New Cancer Immunotherapies New Technique Allows Scientists to Study Parasitic Infections One Cell at a Time Human Textiles to Repair Blood Vessels Cancer-Causing Culprits Will Be Caught by Their DNA Fingerprints Could Resetting Our Internal Clocks Help Control Diabetes? Movement Study Could Be Significant in Helping Understand Brain Rehabilitation Understanding How Cells Defend Their Genome Against Invaders Researchers Discover New Piece of the Puzzle for Parkinson's Disease Never Too Late to Quit: Protective Cells Could Cut Risk of Lung Cancer for Ex-Smokers New Gene Correction Therapy for Duchenne Muscular Dystrophy Can Lithium Halt Progression of Alzheimer's Disease? A New Blood Component Revealed 'Ancient' Cellular Discovery Key to New Cancer Therapies Human Exposure to Aluminum Linked to Familial Alzheimer's Disease Cheap Drug May Alleviate Treatment-Resistance in Leukemia Study Uses Eye Movement Test to Confirm Brain Aging Effects Physicists Design 'Super-Human' Red Blood Cells to Deliver Drugs to Specific Targets Artificial Muscle Sheets Transform Stem Cells into Bone Cell 'Hands' to Unlock Doors in Health Research, Drug Design, and Bioengineering Gene Therapy Shown to Offer Long-Term Benefits for People with Hemophilia A Possible Dementia Vaccine Closer After Mice Studies Half the Amount of Chemo Prevents Testicular Cancer from Coming Back, New Trial Shows Synthetic Nanopores Made from DNA Finding a Non-Invasive Way to Predict Effectiveness of Cancer Therapy How a Protein in Your Brain Could Protect Against Alzheimer's Disease RNA Modification: Methylation and Mopping Up Deeper Understanding of Irregular Heartbeat May Lead to More Effective Treatment Scientists Uncover Structure of Key Pneumonia Virus Enzyme Identification of a Key Protein Linked to Aging 'Junk DNA' Affects Inherited Cancer Risk Long-Term Study Finds Faster Breast Cancer Radiation Treatment as Effective as Long Course How Stem Cells Decide Their Identity New Approach to Treating Cystic Fibrosis Could Lower Risk of Lung Transplants and Death Micro Implants Could Restore Standing and Walking Encouraging Normal Liver Cells to Fight Cancer Using Gene Scissors to Detect Diseases Insights into Versatile Molecular Death Switch Cutting Neurodegeneration Associated with Dementia, in Animal Models Scientists Discover How the Molecule-Sorting Station in Our Cells Is Formed and Maintained Bacterial Protein Impairs Important Cellular Processes RNA Regulation Is Crucial for Embryonic Stem Cell Differentiation Scientists Close In on Malaria Vaccine Complex Organ Models Grown in the Lab Gut Microbiota Imbalance Promotes the Onset of Colorectal Cancer Researchers Find New Way to Target Childhood Cancer Small RNAs Link Immune System and Brain Cells Protein Could Offer Therapeutic Target for Pancreatic Cancer Oxygen Deficiency Rewires Mitochondria Any Amount of Running Linked to Significantly Lower Risk of Early Death Milk from Teeth: Dental Stem Cells Can Generate Milk-Producing Cells A Protein that Pulls the Brake on Nerve Growth Pitfalls with Nanocontainers for Drug Delivery New Gene Therapy for Epilepsy Provides On-Demand Release of Endogenous Substance New Method Identifies Aggressive Breast Cancer Protein in Blood Protects Against Neuronal Damage After Brain Hemorrhage Machine Learning Tool Refines Personalized Immunotherapy Tissue Damage Caused by a Heart Attack to Be Reduced by 30%? Changes Associated with Alzheimer's Disease Detectable in Blood Samples Habitual Tea Drinking Modulates Brain Efficiency: Evidence from Brain Connectivity Evaluation Whole Genome Sequencing Could Enable Personalized Cancer Treatment Simple Cardiac Risk Score Can Predict Problems with Blood Flow in the Brain Cannabis Study Reveals How CBD Offsets the Psychiatric Side-Effects of THC Fruit Flies Live Longer with Combination Drug Treatment Tiny, Biocompatible Nanolaser Could Function Inside Living Tissues Antimicrobial Resistance Is Drastically Rising Discovery of Novel Cancer Signaling Mechanism and Design of New Anticancer Compound Alcohol-Producing Gut Bacteria Could Cause Liver Damage Even in People Who Don't Drink Rethinking How Cholesterol Is Integrated into Cells Same But Different: Unique Cancer Traits Key to Targeted Therapies Alzheimer's Disease Risk Gene APOE4 Impairs Function of Brain Immune Cells Discovery Concerning the Nervous System Overturns a Previous Theory Machine Learning Improves the Diagnosis of Patients with Head and Neck Cancers B Cells Linked to Immunotherapy for Melanoma Breast Cancer Can Form 'Sleeper Cells' After Drug Treatment How Bacteria Behind Hospital Infections Block Out Antibiotics It's Never Too Late to Start Exercising, New Study Shows Scientists Uncover Key New Molecules that Could Help to Tackle Tooth Loss and Regeneration Psychosensory Electronic Skin Technology for Future AI and Humanoid Development Scientists Identify Potential Cause of Statin-Related Muscle Pain Removing Cancer's Protective Barrier Could Boost Immunotherapy Treatments When a Diseased Liver Disrupts the Brain Insomnia Tied to Higher Risk of Heart Disease and Stroke Optic Nerve Stimulation to Aid the Blind Relaxing of Regulations for Regenerative Medicines Has Cascading Effect Internationally 3D Miniature Livers Lead the Way to Patient-Specific Drug Discovery Liver Transplants Could Be Redundant with Discovery of New Liver Cell How HIV Infection May Contribute to Wide-Ranging Metabolic Conditions Beyond Finding A Gene: Same Repeated Stretch of DNA in Three Neurodegenerative Diseases First Impressions Go a Long Way in the Immune System Gut Microbes May Affect the Course of ALS C. Difficile Resists Hospital Disinfectant, Persists on Hospital Gowns, Stainless Steel New Gene Linked to Healthy Aging in Worms Insects Feel Persistent Pain After Injury, Evidence Suggests Pain Signaling in Humans More Rapid than Previously Known World First: Homing Instinct Applied to Stem Cells Show Cells 'Home' to Cardiac Tissue Imprinted Spheres Fight Breast Cancer Researchers Grow Active Mini-Brain Networks Protein Linked to Aggressive Skin Cancer New Basic Understanding of How Lung Cancer Spreads: Antioxidants Implicated Growing Embryonic Tissues on a Chip Artificial Nose Identifies Malignant Tissue in Brain Tumors During Surgery Rheumatoid Arthritic Pain Could Be Caused by Antibodies Molecular Chatter Makes for a 'Hot Tumor' Worm Study Sparks Hope for Slowing Muscle Decline How Genes Interact to Build Tissues and Organisms Alzheimer's Disease Protein Links Plaques to Cell Death in Mice Major Stem Cell Discovery to Boost Research into Development and Regenerative Medicine Coat of Proteins Makes Viruses More Infectious and Links Them to Alzheimer's Disease Virtual Reality Can Spot Navigation Problems in Early Alzheimer's Disease Scientists Discover Signaling Circuit Boards Inside Body's Cells Cancer Cells Are Quick-Change Artists Adapting to Their Environment A Step Closer to Identifying Cause of a Blinding Disease Insulin Under the Influence of Light How Molecular Escorts Help Prevent Cancer Dietary Cholesterol or Egg Consumption Do Not Increase the Risk of Stroke, Finnish Study Finds Why Hodgkin's Lymphoma Cells Grow Uncontrollably How To Starve Triple Negative Breast Cancer How Mutations Lead to Neurodegenerative Disease Alzheimer's Therapy Passes Another Important Test Multiple Sclerosis: Discovery of a Mechanism Responsible for Chronic Inflammation How the Immune System Recognizes Pathogens: Interaction Time Protein Protects Against Non-Alcoholic Fatty Liver Disease Opportunistic Cancer Cells 'Slip Through the Gaps' to Spread Through Blood Vessels A Model to Decipher the Complexity of Gene Regulation Embryo Stem Cells Created from Skin Cells Urine Test Could Prevent Cervical Cancer, Study Finds Key to Targeting Dormant Cancer Cells Lung Cancer Under-Recognized in People Who Have Never Smoked Engineers Make Injectable Tissues a Reality Precise Decoding of Breast Cancer Cells Creates New Option for Treatment Scientists Print First 3D Heart Using Patient's Biological Materials Plant Compounds Could Provide the Answer for Blindness Caused by Diabetes, Say Experts Mapping Cellular Diversity by Looking for Common Topics of Gene Control Genes Linked with the Most Common Liver Cancer Scientists Rejuvenate Stem Cells in the Aging Brain of Mice Harnessing Photonics for At-Home Disease Detection Global Microbial Signatures for Colorectal Cancer The Inflammation Connection A Cellular Protein as a 'Gas Pump Attendant' of Cancer Development Artificial Intelligence Can Predict Premature Death, Study Finds Tissue Engineering: Hydrogel for Enhanced Cell Encapsulation and Delivery Engineering Cellular Function Without Living Cells New Heart Valve Aimed at High-Risk Patients Genetic Variants May Influence Poststroke Recovery Oscillation in Muscle Tissue Smoking, High Blood Pressure, Diabetes and Obesity Each Linked to Unhealthy Brains New 'Tracers' Improve Diagnosis of Cancer and May Be Useful for Treatment When a Cell's 'Fingerprint' Can Be a Weapon Against Cancer Sinister Blastocystis: A Clandestine Killer of Good Bacteria Revealed Precision Drugs Could Unmask Cancers to Immune System and Boost Effects of Immunotherapy Plasma Protein May Hold Promise for Wound Scaffolds Discovery of the Genetic 'Conductor' of Brain Stem Cells New Quantum Sensor Could Improve Cancer Treatment Mini Kidneys from Urine Cells A Common Genetic Signature Has Been Discovered Among Three Cancer Prone Rare Skin Diseases New Protein 'Switch' Could Be Key to Controlling Blood-Poisoning and Preventing Death from Sepsis Researchers Discover 'Chromosome Scanner' that Protects Against Cancer A New Sequencing Method to Detect DNA Modifications of Relevance to Cancer Immune System's Unknown Messenger New Form of Hereditary Osteoporosis Keeping Active in Middle Age May Be Tied to Lower Risk of Dementia Increased Muscle Mass Improves Response to Cancer Treatment First Model of Mitochondrial Epilepsy The Lypla1 Gene Impacts Obesity in a Gender-Specific Manner Blood Clot Discovery Could Pave Way for Treatment of Blood Diseases Human Cells Can Change Job to Fight Diabetes New Therapy for Aggressive Blood Cancer Discovered When Good Cells Go Bad: Regulating the MS-Causing Properties of Th17 Cells Shared Genetic Marker Offers New Promise in Targeting Specific Ovarian and Lung Cancers Researchers Are Worried that People with Chronic Disease Are Not Being Active Enough Newly Discovered Immune Cells Play Role in Inflammatory Brain Diseases More Than 100 New Gut Bacteria Discovered in Human Microbiome It's All in the Code: Protein Production Efficiency Can Be Predicted by Gene Sequence Trial Using Donated Eye Tissue Offers Stem Cell Surgery Hope Cancer: A Mutation That Breaks Gene Interplay in 3D Key Protein in the Production of Insulin Discovered Identical Twins Light the Way for New Genetic Cause of Arthritis Genetic Study Reveals Possible New Routes to Treating Osteoarthritis Early Prediction of Alzheimer's Progression: Blood Protein Gene Therapy Promotes Nerve Regeneration New Therapeutic Avenue in the Fight Against Chronic Liver Disease Synaptic Logic for Connections Between Two Brain Hemispheres Engineered T Cells Promote Long-Term Organ Transplant Acceptance Mesenchymal Stems Cells: Vital Role in Repair of Damaged Organs How Herpes Viruses Shape the Immune System Researchers Uncover New Mechanism of Gene Regulation Involved in Tumor Progression New Materials Could 'Drive Wound Healing' By Harnessing Natural Healing Methods Our Bodies May Cure Themselves of Diabetes in the Future A Boundary Dance of Amyloid-Beta Stepping into Dementia Bacteria Found in Ancient Irish Soil Halts Growth of Superbugs: New Hope for Tackling Antibiotic Resistance Breast Cancer Drugs Could Help Treat Resistant Lung Cancers Molecule Discovery Holds Promise for Gene Therapies for Psoriasis Giving Birth Associated with 14 Percent Higher Risk of Heart Disease and Stroke New Genetic Testing Technology Enhances Precision of Analysis of Clinical Biomarkers Communication Between Neural Networks Biologists Identify Promising Drug for ALS Treatment Discovery of New Neural Mechanism Underlying Anxiety Points to Possible Treatments Lethal Combination: Drug Cocktail Turns Off the Juice to Cancer Cells New Light-Based Technology Reveals How Cells Communicate in Human Disease First Baby Born via Uterus Transplant from a Deceased Donor Underlying Mechanisms of 3D Tissue Formation Cutting the Legs Off Cancer Scientists Develop 'Contact Lens' Patch to Treat Eye Diseases A Molecule for Fighting Muscular Paralysis Hidden Estrogen Receptors in Breast Epithelium How Your Muscles Form How Pneumococci Challenge Our Immune System Sitting Is NOT the New Smoking, Contrary to Popular Myth Breakthrough for Treatment of Fibrotic Diseases Scientists Bring New Hope to Brain Tumor Patients Scientists Figure Out How to Measure Electrical Activity in a Fetal Heart Cooling 'Brains on Fire' to Treat Parkinson's Nerve-On-A-Chip Platform Makes Neuroprosthetics More Effective New Human Cell Structure Discovered AI Doctor Could Boost Chance of Survival for Sepsis Patients Losing Control of Gene Activity in Alzheimer's Disease Cells that Change Jobs to Fight Diabetes Antibodies Linked to Heart Attacks New Reservoir Computer Marks First-Ever Microelectromechanical Neural Network Application Scientists Create Synthetic Prototissue Capable of Synchronized Beating New Function of a Key Component in the Immune System Discovered Protein Dynamics: Molecular Machines at Work AI Could Predict Cognitive Decline Leading to Alzheimer's Disease in the Next Five Years Stem Cells Organize Themselves into Pseudo-Embryos Dutch Study Estimates 1 in 2 Women and 1 in 3 Men Set to Develop Dementia/ Parkinsonism/Stroke New Knowledge on How Neurons Talk to Muscles Immune System's Balancing Act Keeps Bowel Disease in Check Combination Antibody Therapy Results in Long-Term Viral Suppression in HIV Infection Molecule Capable of Halting and Reverting Parkinson's Neurodegeneration Identified Tumor Cell Expansion Challenges Current Physics Lung Cancer Drug Could Be Repurposed to Target 'Zombie' Proteins Linked to Leukemia Deciphering the Link Between Skin Allergies and the Gut Microbiota Drugs That Stop Mosquitoes Catching Malaria Could Help Eradicate the Disease First Gut Bacteria May Have Lasting Effect on Ability to Fight Chronic Diseases New Blood Test Detects Early Stage Pancreatic Cancer Patients with Sepsis at Higher Risk of Stroke, Heart Attack After Hospital Discharge New Technique Reveals How Zika Virus Interacts Inside Our Cells Bioadhesive, Wirelessly-Powered Implant Emitting Light to Kill Cancer Cells Scientists Developing New Blood Test to Screen for Secondary Heart Attack Breaking Osteoporosis: New Mechanism Activates Bone-Building Cells Superbug Discovery Renews Hope for Antibiotic Treatment Attention Network Plays Key Role in Restoring Vision After Brain Damage Anti-Inflammatory Use During Surgery Could Improve Cancer Outcomes Like A Zipper -- How Cells Form New Blood Vessels Drug Could Aid Recovery After A Heart Attack Epigenetic Analysis of Aggressive Brain Tumors New Immunotherapy Inhibits Tumor Growth and Protects Against Metastases Acute Leukemia: Drug Trials Show 50 Percent Cure Rate in Lab Mice New ESMO Tumor DNA Scale Helps Match Patients with Cancer to Optimal Targeted Medicines Powerful Molecules Provide New Findings About Huntington's Disease Switching DNA and RNA On and Off Stress Hormone Is Key Factor in Failure of Immune System to Prevent Leukemia Bacteria-Fighting Polymers Created with Light Scientists Propose a New Lead for Alzheimer's Research Chemicals Found in Vegetables Prevent Colon Cancer in Mice Blood Test Could Detect Kidney Cancer Up To 5 Years Earlier Amputees Feel as Though Their Prosthetic Limb Belongs to Their Own Body PET Tracer Identifies Estrogen Receptor Expression Differences in Breast Cancer Patients Why Weight Loss Produces Remission of Type 2 Diabetes in Some Patients Research into Cell-To-Cell Signaling Mechanism May Lead to New Cancer Treatments Regenerative Medicine: Cell Conversion Factors Predicted A Reliable, Easy-to-Use Pre-Clinical Model for Investigating Bone Metastasis A Brain Injury Diagnosed with a Single Drop of Blood Gene Study Pinpoints Superbug Link Between People and Animals Scientists Generate Key Life Event in Artificial Mouse 'Embryo' Created from Stem Cells Scientists Uncover New Connection Between Smell and Memory Turmeric-Derived Eye Drops Could Treat Glaucoma: Study Blindness Gene Discovered Ultra-Thin Sensor Makes Inflammation Testing and Curing 30 Times Faster New Method for Turning Skin Cells into Pluripotent Stem Cells New Hope for Patients with Incurable and Disabling Hand Condition, Dupuytren's Disease Melting Bacteria to Decipher Antibiotic Resistance New Drug and Material Discoveries to Be Untangled in VR Direct Conversion of Non-Neuronal Cells into Nerve Cells Bacteria Can 'Divide and Conquer' to Vanquish Their Enemies New Sensor Technology Enables Super-Sensitive Live Monitoring of Human Biomolecules Important Step Towards A Computer Model That Predicts the Outcome of Eye Diseases Challenging Our Understanding of How Platelets Are Made Dementia Risk Increased In 50-Year-Olds with Blood Pressure Below Hypertension Threshold Protective Mechanism Against Atherosclerosis Discovered Discovery Shines Light on the Mystery of Cell Death in MS Smoking and Diabetes Linked to Brain Calcifications Scientists Unravel Molecular Mechanisms of Parkinson's Disease What Effect Does Transcranial Magnetic Stimulation Have on the Brain? New Tool Enables Big-Scale Analysis of Single Cells New Theory on Why More Women Than Men Develop Autoimmune Diseases Scientist Reveals Likely Cause of Childhood Leukemia Deadly Malaria's Evolution Revealed Daily Egg Consumption May Reduce Cardiovascular Disease Flowing Cells in a Wavy Microchannel for Effective Size-Based Cell Sorting Expert Consensus Finds That Higher Protein Intake Benefits Adult Bone Health 3D Images of Cancer Cells in the Body IgG Antibodies Activate Blood Platelets and Contribute to the Severity of Anaphylaxis 'Universal Antibodies' Disarm Various Pathogens A New Mechanism for Neurodegeneration in a Form of Dementia Neuro Filter Sharpens Visual Processing Tracing the Footprints of a Tumor: Genomic 'Scars' Allow Cancer Profiling Our Vulnerable Nervous System: What Affects Its Protective Sheaths? Scientists Unearth Vital Link Between Fat, Immunity and Heat Regulation A Non-Coding RNA Lasso Catches Proteins in Breast Cancer Cells Technology Used to Map Mars Now Measuring Effect of Treatment on Tumors What If You Could Know That Your Mild Cognitive Impairment Wouldn't Progress Stem Cells from Adults Function Just as Well as Those from Embryos Commonly Prescribed Heartburn Drug Linked to Pneumonia in Older Adults Labeling and Detecting RNA Modifications Enigmatic Gene Critical for A Healthy Brain Three-Fold Higher Risk of Cancer After Acute Thrombosis in the Leg Antiepileptic Drugs Increase Risk of Alzheimer's and Dementia Blood Flow Is a Major Influence on Tumor Cell Metastasis Tracing the Origin of Each Cell in a Zebrafish The Absence of a Single Mitochondrial Protein Causes Severe Inflammation Digital Penicillin Production Solution To 50-Year-Old Mystery Could Lead to Gene Therapy for Common Blood Disorders Novel PET Imaging Agent Could Help Guide Therapy for Brain Diseases First Proof a Synthesized Antibiotic Is Capable of Treating Superbugs Brain Development Disorders in Children Linked to Common Environmental Toxin Exposures Captured on Film for the First Time: Microglia Nibbling on Brain Synapses Breast Cancer: New Mechanism Discovered for Dissemination of Metastatic Tumor Cells How a Genetic Mutation Can Interfere with the Powerhouses of Cells Staphylococcus Aureus: A New Mechanism Involved in Virulence and Antibiotic Resistance Metastatic Cancer Cells Modify Bone Remodeling with Small RNA Secretion in Bone Metastasis Trial Shows Safety of Drugs for Irregular Heartbeat Patients Undergoing Treatment New Method Manages and Stores Data from Millions of Nerve Cells -- In Real Time New Osteoarthritis Genes Discovered 'Digistain' Technology Offers Revolution in Detailed Cancer Diagnosis Scientists Find Treasure Trove Of 110 Genes Linked to Breast Cancer Calcium May Play a Role in the Development of Parkinson's Disease How the Insulin Receptor Works D-Galactose Affects Ageing Male and Female Brains Differently Electrical Implant Reduces 'Invisible' Symptoms of Man's Spinal Cord Injury Genes Activated in Metastasis Also Drive the First Stages of Tumor Growth Microanalysis of Biological Samples for Early Disease Detection Very Long-Chain Lipids Could Help Prevent Dry Eye Disease Ribosomes Found to Induce Somatic Cell Pluripotency Diabetes Doubles Chance of Developing Cataract Fiber Optic Sensors That Dissolve in the Body Created Low Muscle Strength Identified as Early Risk Factor for ALS Wireless Light Switch for Targeted Cancer Therapy Researchers Map Out the Atlas of Gene Regulators in Human Cardiac Cells for the First Time Hidden Genetic Effects Behind Immune Diseases May Be Missed, Study Suggests Vitamin Deficiency 'Puts Cancer Cells into Hibernation' The Sugar-Attaching Enzyme That Defines Colon Cancer How Treating Eczema Could Also Alleviate Asthma First Vaccine in the World Developed Against Grass Pollen Allergy Blasting Dental Plaque with Microbubbles Multi-Gene Test Predicts Early Heart Disease Risk Mechanism That Converts White Fat to Brown Identified An Organ-On-A-Chip Device That Models Heart Disease Novel Nanomedicine Inhibits Progression of Pancreatic Cancer in Mice Novel Mechanism That Protects from Glioblastoma Identified From the Omelet to the Egg: Reversing Protein Aggregations Using Viruses to Fight Viruses: New Approach Eliminates 'Dormant' HIV-Infected Cells Scientists Discover Blood Sample Detection Method for Multiple Sclerosis Yeast Can Be Engineered to Create Protein Pharmaceuticals How to Tackle Drug Resistant Parasites That Cause Killer Disease Malaria Liver Cancer: Lipid Synthesis Promotes Tumor Formation Long-Term Prevention of Organ Rejection Researchers Fold a Protein Within a Protein Tapeworm Drug Fights Prostate Cancer Deadly Combination in Neurodegenerative Diseases Revealed Epigenetic Editing Reveals Surprising Insights into Early Breast Cancer Development Engineering Non-Immune Cells to Kill Cancer Cells Effective Therapy Against Glioblastoma by Attacking Telomeres Misregulated Protein Breakdown Promotes Leukemias and Brain Cancer Boy Is Given New Skin Thanks to Gene Therapy Virtual Reality Reduces Phantom Pain in Paraplegics Important Mechanism of Epigenetic Gene Regulation Identified New Self-Regulating Nanoparticles Could Treat Cancer Five New Malaria Targets That Could Lead to an Effective Vaccine Transforming Prostate Cancer Treatment Pathway By Which Blood Cells Release a Potent Signaling Factor Discovered Cancer: New Compound Targets Energy Generation, Thereby Killing Metastatic Cells No Evidence That Widely Marketed Technique to Treat Leaky Bladder/Prolapse Works New Antibiotic Resistance Genes Found Bolstering Fat Cells Offers Potential New Leukemia Treatment Wound Care: Patch Could Improve Healing and Reduce Scarring Regenerating Tissues with Gene-Targeting Molecules Drug Targeting Technique Could Aid Therapies for Immune Diseases Cells Programmed Like Computers to Fight Disease Antimalarial Drug Combined with Light Sensitive Molecules for Promising Treatment of Cancer Why We Did Not Evolve to Live Forever: Unveiling the Mystery of Why We Age Cancer Drug Stimulates Tripolar Mode of Mitosis Researchers Discover New, Abundant Enzyme That Helps Bacteria Infect Animals Patients to Benefit from New 3-D Visualizations of the Heart New Mindset in the Search for Stroke Therapies Asthma Medicine Halves Risk of Parkinson's Newly Emerged Superbug Discovered New Immunotherapy Combination Effective at Killing Cancer Cells Worm Infection Reveals Cross-Talk in the Lymph Nodes Eating Triggers Endorphin Release in the Brain Brain's Defense Cells Live Longer Than Expected New Light on Link Between Gut Bacteria and Anxiety The Pancreas Provides a Potential Drug Candidate for Brain Disease Trigeminal Nerve Stimulation Shows Promise for Management of Traumatic Brain Injury How Human Cells Maintain the Correct Number of Chromosomes Dementia: BACE Inhibitor Improves Brain Function New Surgical Strategy Offers Hope for Repairing Spinal Injuries Blocking the Back-Door that Cancer Cells Use to Escape Death by Radiotherapy Understanding Cell Segregation Mechanisms Which Help Prevent Cancer Spread Clues to Healing Spinal Cord Injuries Scientists Enlist Baker's Yeast in a Hunt for New Medicines Fewer Infections in Mechanical Heart Valves Adrenal Gland Activity Changes Under Severe Stress Causing Abnormal Release of Glucocorticoid Stress Antibiotic Insight May Help in Battle Against Bacterial Resistance Two New Genes Linked to Alzheimer's Risk Tracking the Mechanisms of Artery Formation Artificial Sweeteners Linked to Risk of Weight Gain, Heart Disease, and Other Health Issues New Hope for Neuro-Tumor Patients High-Speed Whole-Brain Imaging Improves Understanding of Brain Disease in Animals and Humans Killing Cancer in the Heat of the Moment Fasting Blood Sugar, Fasting Insulin Identified as New Biomarkers for Weight Loss Insulin Release Is Controlled by the Amount of Epac2A at the Secretory Vesicles Bowel Cancer Diagnosis Delayed by Other Illness Vegetable Coloring Agent Lutein May Suppress Inflammation Watching Cancer Spread in Pre-Clinical Trials Genetic DJ: Growing Cells Remix Their Genes Traumatic Brain Injury Associated with Dementia in Working-Age Adults Neuron-Integrated Nanotubes to Repair Nerve Fibers Cancer Hijacks Natural Cell Process to Survive Existing Drugs Could Benefit Patients with Bone Cancer, Genetic Study Suggests Distant Brain Regions Selectively Recruit Stem Cells Viral Vectors for Gene Transfer Travel Longer Distances in the Brain Than Thought Novel Approach May Improve Valve Function in Some Patients Cardiac Stem Cells from Heart Disease Patients May Be Harmful New Genetic Technique Could Help Identify Potential Drug Targets for Malaria When Healthy Cells Stimulate the Migration of Tumor Cells Anorexia Nervosa Has a Genetic Basis Splitting Cells: How a Dynamic Protein Machinery Executes 'The Final Cut' Vitamin C and Antibiotics: A New One-Two 'Punch' for Knocking-Out Cancer Stem Cells Analysis of Complex Protein Interactions Subduing the Rebellion: Unmasking Rogue Cells in the Immune System Inroads Made into Finding Out How T-Cell Acute Lymphoblastic Leukemia Develops Unexpected Presence of Glucose Receptor in Ovarian Cancer Links Metabolism to Most Aggressive Cases New 'GPS' Neuron Discovered Childhood Obesity Causes Lasting Damage to the Body The Immune System Promotes Spontaneous Heart Regeneration How Cancer Cells Flood the Lung New Hope for Slow-Healing Wounds Disrupted Fat Breakdown in the Brain Makes Mice Dumb Epigenetic Program Leading to Vessel Differentiation New Molecular Target to Improve Neuroblastoma Treatment New Ways to Detect and Monitor Inflammatory Bowel Disease Potential Risks of Common MS Treatment Revolutionary Eye Drops to Treat Age-Related Blindness Virus Study Targets Infection Linked to Birth Defects Hepatitis C Virus: Possible New Approach to Battling Infections New Cell Separator with Humble Beginnings Could Revolutionize Medical Advances in Cancer and Alzheimers Research New Roadmap Provides Blueprint to Tackle Burden of Asthma DNA Fingerprinting Reveals How Malaria Hides from Our Immune System Biochemical Pathways of Kidney Disease Revealed Stroke Prevention May Also Reduce Dementia MicroRNA Reduces Stroke Risk Metastatic Breast Cancers: Characterizing the Profile of Metastases for Improved Treatment Novel Molecular Pathway in Aggressive Breast Cancer Offers Potential Therapeutic Targets Solving a Cell Membrane Mystery Ultrasensitive Detection of Protein Linked to Multiple Autoimmune Diseases Tuberculosis: Researchers Uncover How Bacteria Burst Our Cells Cause of An Inherited Neurological Disorder Discovered New Mouse Developed That Could Provide Advance Warning of Next Flu Pandemic Early Life Antibiotic Use Linked to Inflammatory Gut Diseases in Adulthood Preclinical Research: More Reliably Predicting What Will Work Cortisol Excess Hits Natural DNA Process and Mental Health Hard Dementia: The Right to Rehabilitation Why Does the Same Exercise Exert Effects on Individuals Differently? Interferon-Beta Producing Stem Cell-Derived Immune Cell Therapy on Liver Cancer Novel Drug Delivery Beats Swine Flu at Gene Level Link Between Vitamin D Treatment and Autism Prevention New Role for Immune Cells in Preventing Diabetes and Hypertension New Toxic Pathway Identified for Protein Aggregates in Neurodegenerative Disease New 'Gene Silencer' Drug Reduce Cholesterol by Over 50 Percent How Gut Inflammation Drives the Evolution of Harmful Bacteria Prostate Cancer Trial Shows Treating with Precision Radiotherapy Cuts Course of Treatment by 50 Percent Effects of Gut Flora Revisited Intensive Medical Treatment Can Reverse Type 2 Diabetes Shedding New Light on Immune System Response to Zika Virus Role of Rogue Protein PAK4 Confirmed in Pancreatic Cancer Cells Tumor Suppressor Promotes Some Acute Myeloid Leukemias, Study Reveals 'Anti-Aging' Hormone Could Unlock New Treatments for Kidney and Heart Disease Nicotine Exposure During and After Pregnancy Can Cause Hearing Problems in Children Inducing an Identity Crisis in Liver Cells May Help Diabetics Organo-Metal Compound Seen Killing Cancer Cells from Inside Transporter of Thyroid Hormones Is Crucial for the Embryonal Development of the Brain Hard Shell, Healthy Kernel: Nuts Can Inhibit the Growth of Cancer Cells Ground-Breaking Method for Screening the Most Useful Nanoparticles for Medicine Potential New Cancer Treatment Activates Cancer-Engulfing Cells Single-Molecule Technique to Explain Thermodynamic Behavior of Complex Biomolecular Systems 'Survival Gene' Stops Strains of TB Mutating into Deadly 'Superbugs' Predicting and Preventing Prostate Cancer Spread Scientists Decipher a Mechanism in Serious Skin Infections Personalized Treatment for Heart Failure Ion Treatments for Cardiac Arrhythmia Network of Molecular Interactions in Brain Cells Infected by Zika Virus Reveals New Therapeutic Targets 3D Bioprinter to Print Human Skin World Still 'Grossly Underprepared' for Infectious Disease Outbreaks A New Index for the Diagnosis of Non-Alcoholic Fatty Liver Disease Targeted Therapy for Sleep Disorders Helps Patients with Muscular Dystrophy Reducing the Radioresistance of Cancer Sensory Stimuli Control Dopamine in the Brain Bloodstream Infections: Most Common Type of Health Care-Associated Infections in Children Dengue, Zika Virus Family Uses an Unexpected Approach to Hijack Human Cell Machinery Autoimmunity and Infections: When the Body Fights Itself New Research Describes How Bacteria Resists 'Last-Resort' Antibiotic New Treatment for a Rare Form of Encephalitis Lung-Specific Ebola Infection Found in Recovering Patient How Bacteria Survive Antibiotic Treatment Important Element of Immune Defense Against Fungal Infections Discovered Brain Shrinkage in Multiple Sclerosis Associated with Leaked Protein in Blood Therapy Response in Brain Tumor Cells Is Linked to Disease Prognosis New Drug Limits and Then Repairs Brain Damage in Stroke New Target Receptor Discovered in the Fight Against Obesity World of Viruses Uncovered Mechanism Revealed for Side Effects of Drug Used in Hematopoietic Stem Cell Harvesting Scientists Step Closer to Halting Spread of Lung Cancer Connection Found Between Memory Mechanisms, Resistance to Epilepsy Depression in Young People Affects the Stomach, Anxiety the Skin Imaging Technique Measures Toxicity of Alzheimer's, Parkinson's Proteins Only Half of a Chromosome Is DNA, 3-D Imaging Reveals Blood Test Could Predict Best Treatment for Lung Cancer Reconditioning the Brain to Overcome Fear Genetic Mutations That Lead to Macular Degeneration Blindness Mapped By New Research Genes and Age Determine Susceptibility to Lyme Disease Researchers Find Immunotherapy Treatments Better for Advanced Skin Cancer Multidrug-Resistant Bacteria from Chickens Pose Risk to Human Health Faster Diagnosis of Sepsis Pathogens Vitamins A and C Help Erase Cell Memory Substance with the Potential to Postpone Aging Soft Robots That Mimic Human Muscles Scientists Find New Metabolic Pathways to Resist Viruses New Antibody Therapy Permanently Blocks HIV-Like SIV Infection in Monkeys High Cholesterol Triggers Mitochondrial Oxidative Stress Leading to Osteoarthritis Fungal Infection That Could Help Understand Some Allergies Drivers of Evolution Hidden in Plain Sight Intestinal Diversity Protects Against Asthma New Immune Cell Subset Associated with Progression to Type 1 Diabetes Eating Oats Can Lower Cholesterol as Measured By a Variety of Markers, Review Suggests Are Natural Alternatives to Estrogen Replacement Therapy Safe? Food-Poisoning Bacteria May Be Behind Crohn's Disease Research Uncovers a New Disruption at the Root of Parkinson's Disease Researchers Activate Repair Program for Nerve Fibers Certain Alternative Therapies May Help Patients with Bowel Disorders Keeping Your Synapses Sharp: How Spermidine Reverses Age-Related Memory Decline Brain Diseases Manifest in the Retina of the Eye IL-34: New, Possible Target to Tackle Drug-Resistant Cancer Cells Respiratory Tract Bacterium Uncovered as Trigger for Serious Nervous System Disease New Imaging Technique in Alzheimers Disease Opens Up Possibilities for New Drug Development New Weapon for Hard-To-Treat Bacterial Infections How Cancers 'Invisibility Cloak' Works Custom-Tailored Strategy Against Glioblastomas Multifaceted Genetic Impact of Training Does Physical Activity Lower the Risk of Bacterial Infections? Landmark Map Reveals the Genetic Wiring of Cellular Life Artificial Intelligence Reveals Mechanism Behind Brain Tumor Role Identified for Key Protein in Regeneration of Damaged Newt Retinas Targeting Dormant HIV Memory of a Heart Attack Is Stored in Our Genes Midlife Physical Activity Is Associated with Better Cognition in Old Age Synaptic Connectivity Motifs Contribute to Memory Storage and Retrieval in Hippocampus New Strategy May Help Prevent Kidney Failure in Patients with Diabetes Measuring Forces in the DNA Molecule Aberrant Epigenetic Regulation Behind the Intestinal Symptoms in Celiac Disease Discovery Contributes to Better Understanding of Acquired Resistance to Combination Drug Treatment in Cancer Gut Bacteria Affect Immune Recovery in HIV Patients, Study Finds Gene Defect as a Potential Gateway for Targeted Prostate Cancer Therapy Whole Brain Radiotherapy Offers Little Benefit to People Whose Lung Cancer Has Spread to the Brain, Despite Its Widespread Use New Hope for New Crohn's Disease Treatment Parkinson's Study Could Pave Way for Early Detection Test First Epigenetic Test to Diagnose Tumors of Unknown Origin New Research Reveals Cancers Need a 'Perfect Storm' of Conditions to Develop Elongation by Contraction: Pulling Forces Drive Changes in Cell Shape The First Stage of the Cascade Vertebroplasty Helps Reduce Acute Pain Among Patients With Spinal Fractures From Pollution to Caffeine Intake: Researcher Reveals Dementia Risks Legions of Nanorobots Target Cancerous Tumors with Precision Brain Study Confirms Gene Mutation Link to Psychiatric Disorders New Research Sheds Light on the Role of Proteins and How Synapses Work Active Hedgehog Signaling in Connective Tissue Cells Protects Against Colon Cancer Olfactory Receptors Discovered in Bronchi Dubious Marketing of Stem Cell Therapies Revealed Breakthrough in Understanding How Stem Cells Become Specialized Cerebrospinal Fluid Signals Control the Behavior of Stem Cells in the Brain Machinery Neurons Used to Form and Maintain Their Neuronal Extensions Discovered Modifications to Protein RUNX3 May Promote Cancer Growth Rare Fungus Product Reduces Resistance to Antibiotics Automatically Identifying Brain Regions How New HIV Drugs Lock Virus in Immaturity Uncovering the Mystery of DNA Replication Novel Discovery Improves Profiling of Acute Myeloid Leukemia Patients for Targeted Therapies Promising New Methods for Early Detection of Alzheimer's Disease New Work Aiming to Stop Diabetes, A Major Global Health Challenge An Epigenetic Mechanism Is Involved in Development of Autoinflammatory Diseases Transplant Drug May Provide Benefits After Spinal Cord Injury Where Did It All Go Wrong? Scientists Identify 'Cell of Origin' in Skin Cancers Unsilencing Silenced Genes by CRISPR/Cas9 New Method Provides Better Information on Gene Expression Cancer-Preventing Protein Finds Its Own Way in Our DNA African Subterranean Animal Exhibits 'Extraordinary' Cancer Resistance Personalized Medicine Will Employ Computer Algorithms New Types of Blood Cancer Discovered in Children Scientists Use Modeling to Show the Role of Metabolism and Signaling in Cancer Metastasis Natural Killer Cells Have a Memory Mediterranean Diet High in Healthy Fat Does Not Lead to Weight Gain, According to Randomized Trial New Model of T Cell Activation Targeting Metals to Fight Pathogenic Bacteria Gut Bacteria May Contribute to Poor Health in Patients with Kidney Disease The Brain Needs Cleaning to Stay Healthy Additional Hormone Measurement Reveals Pregnant Women at High Risk of Preeclampsia Modified Microalgae Converts Sunlight into Valuable Medicine Transition Between Arm, Hand Occurs Thanks to a Genetic Switch New Gene for Familial High Cholesterol Blood Marker Determines Who Will Respond to Colorectal Cancer Drug Study of Glioma Susceptibility in Dogs May Yield Insights for Humans Noninvasive Monitor Assesses Patients' Response to Painful Stimulation During Surgery An Old New Weapon Against Emerging Chikungunya Virus Brain Cells That Aid Appetite Control Identified Fastest-Ever Molecular Imaging Reveals Reaction Crucial for Vision Narrow Band Imaging Can Reduce Recurrence of Bladder Tumors New Gene Testing Technology Finds Cancer Risks 'Hiding in Plain Sight' Fiber Optic Biosensor-Integrated Microfluidic Chip to Detect Glucose Levels Vitamin Stops the Aging Process of Organs Synthetic Derivatives of 'Game-Changing' New Antibiotic Could Hold Key to Tackling Drug Resistant Bacteria Breast Cancer Progression: The Devil Is in the Details Got Good Fat? New Hepatocellular Carcinoma Prognostic Model Improves Prediction of Patient Survival A Fatty Liver May Result in a Broken Heart Expert Panel Evaluates Role of Osteoporosis Medications in Fracture Healing Breakthrough in Vaccine Development Sleep Loss Detrimental to Blood Vessels Researchers Uncover 'Local Heroes' of Immune System Older Adults Need Better Blood Pressure and Cholesterol Control to Prevent Cardiovascular Disease Immune Cells in Organ Cavities Play Essential Role in Fast Tissue Repair Cpf1: CRISPR-Enzyme Scissors Cutting Both RNA and DNA Targeted Missiles Against Aggressive Cancer Cells Clinical Trial for Experimental Ebola Drug Publishes Results Unveiling the Grammar of Biological Cells Study May Have Important Implication for the Treatment of Parkinson Disease Brain Responses Found to Originate from Previously Unknown Source How Bioceramics Could Help Fight Gum Disease Scientists Reveal Endocardial Origin of Liver Vasculature Promising New Method Inhibits TB-Causing Bacteria Number of Adults with Diabetes Reaches 422 Million Worldwide, with Fastest Increases in Low and Middle Income Countries Novel Compounds Could Yield More Effective, Less Toxic Cancer Chemotherapy Brain Study Reveals How Long-Term Memories Are Erased 'Precision Medicine' Brings New Relief for Old Diseases Imitating Movements Could Help Alzheimer's Patients Concern: Diabetes and Rising Global Temperature Handheld Surgical 'Pen' Prints Human Stem Cells Embryo Development: Some Cells Are More Equal Than Others Even at Four-Cell Stage 'Clogged-Up' Immune Cells Help Explain Smoking Risk for TB No Increased Heart Failure with Incretin-Based Drugs New Drug Shows Promise Against Muscle Wasting Disease Cells in Standby Mode Researchers Refute Textbook Knowledge on Macrophage Metabolism in Inflammatory Diseases New Role of Environment in Multiple Sclerosis Revealed Scientists Pinpoint Molecular Signal That Drives and Enables Spinal Cord Repair CPAP May Not Improve Glycemic Control in People with Diabetes Early Detection: Colorectal Cancer Rates Declining in Germany Lack of TRPV2 Impairs Thermogenesis in Mouse Brown Adipose Tissue Researchers Dig Up New Molecular Details on 'The Other Type' of Stem Cells Soap Bubbles for Treating Stenosed Blood Vessels New, Non-Invasive Method Determines Whether a Child Is Celiac or Not With Just a Puncture in the Finger Heart Attacks Could Be Reduced by Rethinking the Way We Prescribe Statins Seismic for the Spine: Vibration Technology Offers Alternative to MRI New Insight Into the Possible Risk Factors Associated with Food Allergies Making Better Enzymes and Protein Drugs New Target for Reducing Nerve Pain Identified Genetic Switch Regulating Satiety and Body Weight Identified Two-Pronged Attack Increases Potency of New Anti-Cancer Drugs 'Neoehrlichiosis' Internationally Diagnosed, Successfully Treated in Patients Without Immunodeficiency for the First Time Nanotechnology Delivery System Offers New Approach to Skin Disease Therapies Scientists Make Significant Anti-Aging Breakthrough New Research Adds Additional Layer of Complexity to Human Protein Landscape Seizures and Other Extreme Events in the Brain What Makes the Brain Tick So Fast? Stem Cell Technique Makes Sperm in a Dish Cell Insight Offers Clues on Biological Processes Linked to Fertility Location May Be Key to Effectively Controlling Pain Why Is Calcific Tendinitis So Painful? Anxiety Disorder Three Times More Likely Among Older Adults with COPD Biomolecule's Behavior Under Artificial Conditions More Natural Than Expected Scientists Create Imaging 'Toolkit' to Help Identify New Brain Tumor Drug Targets Drug Does Not Significantly Reduce Duration of Mechanical Ventilation for COPD Patients Scientists Have Put a High Precision Blood Assay into a Simple Test Strip Height Influences Risk of Cardiovascular Disease, Diabetes, and Cancer New Piece of the HIV Puzzle Uncovered Effects on HIV and Ebola Using Mathematics to Improve Human Health Only a Minority of Cancer Cells Affect the Growth, Metastasis of Tumors Adenosine Deaminase May Help the Immune System Fight HIV on Its Own Inflammation Attacks Brain's Reward Center Listeria: Hypervirulent Strains with Cerebral and Placental Tropism Assessing Stem Cells: New Biomarker Developed The Harbingers of Aging Super-Sharp Images Through Thin Optical Fibers Vital Clues to Future Cancer Development in Normal Breast Tissue DNA Neurogenesis: Discovery of a New Regulatory Mechanism It's Complicated: Benefits and Toxicity of Anti-Prion Antibodies in the Brain Epigenetic Switch for Obesity Identifying Another Piece in the Parkinson's Disease Pathology Puzzle Germs, Humans, and Numbers: New Estimate Revises Our Microbiome Numbers Downwards Fat Injection for Breast Reconstruction Doesn't Increase Risk of Recurrent Breast Cancer Making Liver Tissue in the Lab for Transplants and Drug Screening Virtual Bone Biopsy Helps Identify Why People with Diabetes Are at Risk of Bone Fractures Synthetic Biologists Use Bacterial Superglue for Faster Vaccine Development New Research on Inflammation and Cancer B12 Deficiency a Concern for Long-Term Care Important Regulator of Immune System Decoded Protein 'Handbrake' Halts Leukemia in Its Tracks A New Method to Improve the Pre-Operative Diagnosis of Ovarian Cancer Based on Ultrasound Bone Marrow Lesions Can Help Predict Rapidly Progressing Joint Disease Studying 'Inflamm-Aging': Monocytes, Cytokines, and Susceptibility to Pneumonia New Tool May Help Predict Patients' Motor Function Recovery After Stroke Long-Term Outcomes of Preventing Premature Menopause During Chemotherapy Innovation Sheds Light on How Genetic Information Travels From Cell's Nucleus Mechanism of an AIDS Vaccine Candidate Filmed In Vivo In Pursuit of the Causes of Cardiac Hypertrophy Targeted Alpha Therapy's Potential to Eliminate HIV-Infected Cells Protein That Boosts Memory Identified Obesity More Dangerous Than Lack of Fitness, New Study Claims A Multitool for Cells: Understanding the Sense of Smell in Cells Physics Sheds Light on Stem Cell-Derived Organoid Growth, Brain Development Sepsis: Cell Therapy to Repair Muscle Long-Term Impairment New Blood Test Could Predict Arthritis Risk Early Heart Disease: Jamming the Signal Imaging Test Detects Aggressive, Treatment-Resistant Cancers Researchers Use Mathematical Models to Study Brain Cells Essential for Correct Functioning of Neurons Cancer Study Identifies Genes That Stop Onset of Leukemia How Killer -T-Cells Migrate Towards Virus-Infected Cells How Fast You Move Can Predict How Healthy You'll Be iPS-Derived Kidney Cells Successfully Connected to Capillary Network Bacterial Protein Can Help Convert Stem Cells Into Neurons Kidney Cells Made from Reprogrammed Stem Cells Lead Scientists to an Accurate Way to Screen for Toxic Compounds Duchenne Muscular Dystrophy Is a Stem Cell Disease Gene Therapy: A Promising Candidate for Cystic Fibrosis Treatment Blood Sample New Way of Detecting Cancer No New Heart Muscle Cells in Mice after the Newborn Period Revealing the Differences Between Subsets of Immune Cells in the Blood Researchers Create a Worm Model to Investigate a Rare Subtype of Blindness Genetically Modified Mice Reveal the Secret to a Painless Life Researchers Identify Biomarker of Early Lung Cancer that May Increase Survival Researchers Unravel Abilities of the Immune System's Giant Cells New Target Could Eliminate Lurking Cancer Stem Cells Discovered: A Cluster of 60 Proteins That Help Cells Move and Feel New Strategy Discovered for Treating Arthritis Functional Human Liver Cells Grown in the Lab Antibody-Drug Compounds and Immunotherapy to Treat Breast Cancer Stem Cell Study Paves the Way for Patient Therapies Circulating Small Cell Lung Cancer Cells Successfully Cultivated for the Very First Time New Method Can Detect Cancer Cells Before They Form New Tumor Competition Between 'Good Bacteria' Important For Healthy Gut Stem-Cell Scientists Redefine How Blood Is Made, Toppling Conventional 'Textbook' View from 1960s Why Some Genes Are Highly Expressed Researchers Uncover Genes That Can Suppress HCV Infection Within the Liver Immune Cells That Fight Obesity Manipulating the Antennae on Cells Promises New Treatments for Osteoarthritis New Article Reviews Latest Advances in Magnetic Particle Tracking in Cell Therapy Worlds First Lab-in-a-Briefcase Cancer-Causing Parasite May Accelerate Wound Healing Probing Cells' Recycling Centers Reveals New Drug Targets New UCL Study Highlights Genes Associated with Schizophrenia and Obesity Helper Cells in the Brain Could Hold the Clue to Motor Neuron Disease Study Could Help Mitigate Effects of Aging on Blood Stem Cells, Promote New Therapies for Anemia Researchers Discover a New Old Protein in B Cells High Cholesterol Linked to Heightened Risk of Tendon Abnormalities and Pain Link Discovered Between Rare Form of Anemia, Cancer Gene Therapy Prevents Neuronal Death, Disease Effects in Experimental Model of PD Glycan Binding Proteins Modify Parasite Infection in Cells of the Heart Muscle Adult High Blood Pressure Risk Identifiable in Childhood Environmental Memories Transmitted From a Father to His Grandchildren Antioxidants Cause Malignant Melanoma to Metastasize Faster Scientists Build a Digital Piece of a Rat's Brain Certain Blood Markers May Indicate Early Signs of Kidney Disease Detecting HIV Diagnostic Antibodies with DNA Nanomachines Team Succeeds in Producing Photoreceptors from Human Embryonic Stem Cells Targeting Protein Used by Tumors to Help Detect Food Supplies Could Restrict Cancerous Cells' Ability to Grow Researchers Identify Cause of Inherited Form of Extreme Nearsightedness Discovering the Brains Memory Switch Fruit Fly Research Reveals Genetic Mechanisms of Dietary Sugar Sensing Studying Cardiac Arrhythmias in Nematodes Cell Marker Enables Prognosis About the Course of Infections Cancer Test Predicts Treatment Outcome Genes That Protect African Children from Developing Malaria Identified New DNA Stain Lights Up Living Cells A Snapshot of Stem Cell Expression Scientists Discover How to Trap Cancer Cells Before They Spread Intratumor Morphological Heterogeneity of Cancer is Not Related to Chromosome Aberrations DNA Repair Mechanism Manipulated to Delete Genes Without Leaving a Scar A Shift in Energy Processing Pathways Occurs in Immune Cells that Tolerate Pathogens Antimicrobial Film for Future Implants Scientists Uncover How Embryonic Stem Cells Inhibit Virus Expression Discovery of a Triple Barrier that Prevents Cells from Becoming Cancerous Ages Apart: Multifaceted Approach Measured How Brain and Liver Age Differently Digital 'Rosetta Stone' Decrypts How Mutations Rewire Cancer Cells Scientists Discover How Cells Overpower Cancer Drug New Leukemia Gene Stops Blood Cells 'Growing Up' Scientists Move Closer to Curing Common Eye Disorder Tumor Suppressor Genes Curb Growth in Neighboring Cells Chemical-Only Cell Reprogramming Transforms Human and Mouse Skin Cells into Neurons Brazilian Wasp Venom Kills Cancer Cells By Opening Them Up Genes Physically Held in Silencing 'Lock-Down' in Embryonic Stem Cells Study Sheds Light on Ways to Improve Embryonic Wound Healing in Humans Biocompatible Scaffold Enables Beating Heart Cells to Snap Together Like Velcro Researchers Mimic Viral Infection in Colon Cancer Stem Cells When Fat Is the Solution: Using Adipose Cells to Attenuate Chagasic Cardiomyopathy Glial Cells Use Lipids to Direct Neuron Organization in the Spinal Cord Parkinson's Disease Brain Cells at Risk of Burnout, Like an Overheating Motor How Do Harmful Chronic Myelogenous Leukemia Stem Cells Obtain Their Nutrients? Blood Vessel Cells Help Tumors Evade the Immune System Study Sheds Light on Helper Cells and Killer Cells Nanoparticle-Mediated Wound Healing in Diabetes iPS Cells Discover Drug Target for Muscle Disease Transplanted Amniotic Stem Cells Offer Hope for Retinal Diseases Getting Into the Groove Helps Stem Cells Switch Into Heart Muscle Cells Researchers Reveal How Migrating Tumor Cells Produce Protein That Helps Set Up Home in Bones Nerve Cells Use Each Other as Maps Hydrogels Help Stem Cells Accelerate Healing New Stem Cell Therapy Counters the Effects of Osteoporosis in Mice Induced Pluripotent Stem Cells Show Promise for Kidney Treatment Fluorescent Material Reveals How Cells Grow Chameleon Proteins Make Individual Cells Visible The Life and Death of Beta Cells What Hundreds of Biomolecules Tell Us About Our Nerve Cells Soft-Tissue Engineering for Hard-Working Cartilage Dr. Jekyll & Mr. Hyde: Telomere and Telomerase in Stem Cells Stem Cell Therapy Shows Promise in Small Clinical Trial for Rare Lung Disease Gene Therapy Gives Long-Term Protection to Photoreceptor Cells Stem Cells Might Heal Damaged Lungs Scientists Gain New Insights Into 'Antenna' of Human Cells Novel Gene Therapy Control System Regulates Expression of Therapeutic Transgenes Cells Help Viruses During Cell Entry Production of iPS Cells: Discovery of the Fifth Element Stem Cell Transplant Alleviates Symptoms in Lupus Animal Models Spray Application of Respiratory Cells Holds Promise for Tissue Engineering New Cell Division Mechanism Discovered Jumping Genes May Drive Esophageal Cancer New Genomic Analysis Identifies Recurrent Fusion Genes in Gastric Cancers Scientists Discover Origins of the Lymphatic System Dental Pulp Stem Cell Transplants Can Contribute to Peripheral Nerve Regeneration Schwann Cells 'Dine In' to Clear Myelin From Injured Nerves Eavesdropping on the Body: New Device Tracks Chemical Signals Within Cells Premature Aging: Scientists Identify and Correct Defects in Diseased Cells Action Spectrum of Sun Skin Damage Documented Why Human Egg Cells Don't Age Well Colon Cancer: Taking a Step Back to Move Forward New Stem Cell Research Uncovers Causes of Spinal Muscular Atrophy Too Exhausted to Fight and to Do Harm Hope for Patients with Chronic Wounds Build-up of DNA Damage in Nerve Cells Can Lead to Dementia Discovery Targets Dormant Bowel Cancer Cells Before They Form Secondary Tumors Double Agent Skin Cells in Fight Against Dengue Cell Skeleton Structure Breakthrough Factors Released Following Stem Cell Transplantation Therapeutically Impact Serious Burns Tool to Analyze Genes According to Their Evolutionary Profiles Olfactory Cells Transplanted to Treat Spinal Cord Injury How Do Neural Cells Respond to Ischemia? First Embryonic Stem Cell Therapy Safety Trial in Asian Patients with Degenerative Eye Diseases Parallel Sequencing of DNA and RNA Provides Insight into Secret World of Cells Self-Assembling Biomaterial Forms Nanostructure Templates for Human Tissue Formation Beyond Genes: Are Centrioles Carriers of Biological Information? Study Shows Feasibility of Using Gene Therapy to Treat Rare Immunodeficiency Syndrome Cancer Gene Unintentionally Ends the Life of Cancer Cells, Turns Off Life Supporting Genes Fruit Fly Studies Shed Light on Adaptability of Nerve Cells Making Gene Therapy Safer Research Reveals New Possibilities for Islet and Stem Cell Transplantation Nondestructive 3-D Imaging of Biological Cells With Sound Pancreatic Cancer Loses Viral Defenses When Talking With Supporting Cells Patient's Own Fat Cells Transplanted to Treat Osteoarthritis Found May Be Effective Gene Therapy Superior to Half-Matched Transplant for 'Bubble Boy Disease' Staining Method Brings the Reconstruction of All Nerve Cells and Their Connections Within Reach Researchers Develop Harmless Artificial Virus for Gene Therapy Spider Web Protein Provides Stable Substrate for Growing Heart Cells Possible Progress Against Parkinson's and Good News for Stem Cell Therapies Migrating Immune Cells Promote Nerve Cell Demise in the Brain Two Different Fat Graft Techniques Have Similar Effects on Facial Skin Controlling Inflammation in Fat Cells to Fight Obesity-Induced Diabetes New Model to Study HIV Latency in Brain Cells Tissue Scaffold Technology Could Help Rebuild Large Organs Restoring Natural Immunity Against Cancers Scientists Map Surface of Immune Cells Large Doses of Antioxidants May Be Harmful to Neuronal Stem Cells Discovering a New Force Driving Cell Contraction during Development and Organogenesis Finding Hope in the Dark New Procedure to Obtain Induced Pluripotent Stem Cells Bringing Genes into Focus Mobilizing the Skins Defense Cells Recovery of Sensory Function by Stem Cell Transplants Reprogramming of DNA Observed in Human Germ Cells for First Time When the Evil Assumes Power: On the Dominance of Stem Cell Mutations in Age Gene Controls Stem Cells During Muscle Regeneration New Study Shows Stem Cell Treatments Promote Faster Healing in Primates with Spinal Cord Injury Nanotechnology Helps Protect Patients from Bone Infection Stem Cell Therapy for Inherited Skin Blistering Blood to Feeling: McMaster Scientists Turn Blood Into Neural Cells Team Develops Method to Better Identify Genes Involved in Diseases Memory Immune Cells That Screen Intruders as They Enter Lymph Nodes Metabolic Imbalance Increases Risk of Respiratory Diseases in Childhood Gene Regulates Healing Process After Myocardial Infarction Detecting Cancer Cells in Blood Can Give an Early Warning of Treatment Failure Researchers Tweak the Immune System to Target Cells Bearing Tumor Antigens Combining Stem-Cell Therapy with Anti-Scarring Agent May Heal Injured Kidneys New Technique to Chart Protein Networks in Living Cells Protein Mimic Shows Promise as Tissue Engineering Glue Stem Cells Show Promise for Reversing Type 2 Diabetes Researchers Find T Cells Target Dengue Virus at the Skin Infection Site Injured Spinal Cord: Regeneration Possible With Epothilone? Media Portray Unrealistic Timelines for Stem Cell Therapies Cancer-Linked Protein Helps Control Fate of Intestinal Stem Cells Engineered Cells Could Help Tackle the Third Most Common Cancer in Chinese Males Autism Genes Found to Be Associated With Brighter Minds Imperial Scientists Develop New Test That Uses Human Stem Cells to Predict Side Effects of Drugs Limited Self-Renewal of Stem Cells in the Brain Healthy-Looking Prostate Cells Mask Cancer-Causing Mutations Neuron Groups, Not Single Cells, Maintain Brain Stability Bacteria 'Hotwire Their Genes' to Fix a Faulty Motor Immune Cells Repurpose a Stem Cell Transcription Factor as a Pathogen Sensor Scientists Discover New Roles for Viral Genes in the Human Genome 'Stem Cell' Test Could Identify Most Aggressive Breast Cancers Scientists Move Closer to Creating Cartilage from Stem Cells Researchers Identify the Mitochondrial 'Shield' That Helps Cancer Cells Survive Stellate Cells in the Liver Control Regeneration and Fibrosis Skeleton of Cells Controls Cell Multiplication Breakthrough in Understanding How Cancer Cells Metastasize Scientists Grow Leg Muscle from Cells in a Dish Obesity Genes Identified by Worldwide Research Team New Brain Mapping Reveals Unknown Cell Types A Good Night's Sleep Keeps Your Stem Cells Young Breakthrough in the Fight Against Blindness Voltage Tester for Beating Cardiac Cells Changing Stem Cell Structure May Help Fight Obesity Gold Nanotubes Launch a Three-Pronged Attack on Cancer Cells Cerebral Palsy - It Can Be in Your Genes Two New Genetic Variants Linked to Breast Cancer Bacteria Protect Intestinal Tumor Model from Being Killed by Immune Cells Scientists Identify New Population of Regulatory T-Cells New Innovative Method for Mechanical Screening of Biological Cells Developed Understanding the Origins and Function of CD14+ Immune Cells Culture Shock -- Are Lab-Grown Cells a Faithful Model for Human Disease? Researchers Question Treatment of Infertility with Stem Cells Activated Immune Cells Indicate a Favorable Prognosis in Colorectal Cancer Tübingen Researchers Develop New Way of Labeling T-Cells Limitations with Carbon Nanotubes in Blood Facing Medical Devices Discovered Researchers Examine How Terpenes Can Inhibit Growth of Liver Cancer Cells New Method for Analyzing RNA Sequence Data Identifies New Subtypes of Cells Synthetic DNA Gel Points the Way to Printing Artificial Organs Preventing Build-Up of Toxic Fouling Films on Medical Devices Reversing Effects of Early Stage Age-Related Macular Degeneration May Be Possible Cooperation Between Cancer Cells Makes Therapies Ineffective, Suggests New Treatment Subpopulation of Innate Immune Cells May Be Primed Against Threats Like Cancer Dragnet for Epilepsy Genes From Stem Cell to Nerve Cell in a Few Weeks A Scalable Platform for Growing Heart Muscle Cells May Lead to Repair of Damaged Heart Cells Protein Induces Self-Destruction in Cancer Cells How to Attack and Paralyze Myeloma Cells Genetic Discovery About Childhood Blindness Paves the Way for New Treatments Blocking STAT3 in Immune System Cells Increases Anti-Tumor Immunity New Stem Cell Technology Reveals Fresh Insights into Motor Neuron Disease Fast Sorting of CD4+ T Cells from Whole Blood Using Glass Microbubbles Proteases Help Nerve Cells to Navigate Protecting Us from Our Cells New Analysis Methodology May Revolutionize Breast Cancer Therapy Scientists Identify "Naïve-Like" Human Stem Cell Blood Vessel Lining Cells Control Metastasis Chronic High Blood Pressure Increases Risk of Glaucoma Bacteria and Immune Cells Forge a Productive Partnership Sorting Bloodborne Cancer Cells to Better Predict Spread of Disease Malaria-Infected Cells May Latch Onto Healthy Blood Cells for Protection How Cells Defend Themselves Against Antibiotics and Cytostatic Agents Unlocking the Secrets of Pulmonary Hypertension Patient Stem Cells Used to Make Dementia-in-a-Dish; Help Identify New Treatment Strategy Scientists Sweep Cells Apart for Use in Medical Research Most Commonly Prescribed Glaucoma Drug Reduces Risk of Vision Loss by More Than 50 Percent Over Two Years Cell Biologists Discover On-Off Switch for Key Stem Cell Gene Fat Cells Reprogrammed to Increase Fat Burning Genes Show the Way to Better Treatment of Hepatitis C Genes Influence Types of Microbes in Human Gut Gold Nanorods Target Cancer Cells Researchers Treat Heart Attacks with New Gene Therapy Based on Telomerase Enzyme Could Trophoblasts Be the Immune Cells of Pregnancy? Neurons Listen to Glia Cells Genes That Cause Pancreatic Cancer Identified by New Tool Teleophthalmology for Screening, Recurrence of Age-Related Macular Degeneration New View on How Nerve Cells Die in Parkinson's Disease DNA Sequencing Helps Identify Genetic Defects in Glaucoma Nanotechnology Against Malaria Parasites Researchers Identify New Targets for Future CLL Therapies Not All Induced Pluripotent Stem Cells Are Made Equal Genes and Environment: Complex Interactions at the Heart of Personalized Medicine A Mechanism That Alters the Regulation of Calcium in Cells Could Be Involved in Neurodegenerative Disease Researchers Describe a Mechanism That Explains How the Breast Cancer Cells Transform Normal Cells into Tumoral Ones Revealed: How Bacteria Drill Into Our Cells and Kill Them Researchers Use Mathematical Models to Predict How Immune Cells Respond to Disease Why Cancer Cells Grow Despite a Lack of Oxygen Immune Cells from the Spleen Found to Control Chronic High Blood Pressure Research Shows Extremely Limited Diversity Among One Group of Immune Cells and Uncovers a New Population Next Generation Biomarker Detects Tumors Cells and Delivers Anti-Cancer Drugs New Laser Therapy Helps Slow Macular Degeneration Mind the Gap: How New Insight into Cells Could Lead to Better Drugs Gene Therapy Provides Safe, Long-Term Relief for Patients with Severe Hemophilia B A New Nanomachine Shows Potential for Light-Selective Gene Therapy Cataloguing 10 Million Human Gut Microbial Genes Largest-Ever Map of the Human Interactome Predicts New Cancer Genes Alzheimer's Disease: Molecular Signals Cause Brain Cells to Switch Into a Hectic State Lung Cancer Can Stay Hidden for Over 20 Years Multiple Neurodevelopmental Disorders Have a Common Molecular Cause Researchers Develop Nanoparticle Eye Drops to Combat Dry Eye Syndrome Mechanism That Repairs Brain After Stroke Discovered Long-Term Treatment Success Using Gene Therapy to Correct a Lethal Metabolic Disorder Study Provides Vital New Insight into How We Produce New Brain Cells Bowel Cancer Risk Reduced by Adopting Multiple Healthy Behaviors Researchers Aiming to Improve Global Standards for Detection of Leading Cause of Blindness Geneticists Solve 40-Year-Old Dilemma to Explain Why Duplicate Genes Remain in the Genome Superior Hybrid Gold Nanoparticle Cardiac Patch for Tissue Engineering Research Mimics Brain Cells to Boost Memory Power Promising Drug Candidate for Alzheimer's Found in Turmeric Compound Genes Causing Pediatric Glaucoma Contribute to Future Stroke A Second Look at Glaucoma Surgery New Molecule Allows for Increase in Stem Cell Transplants Researchers Discover What Lies Behind the Death of Stem Cells Zebrafish Genes Linked to Human Respiratory Diseases Scientists Reset Human Stem Cells to Earliest Developmental State Twins Help Unravel Glaucoma Genes Five Genes to Predict Colorectal Cancer Relapses Scientists Create Therapy-Grade Stem Cells Using New Cocktail to Reprogram Adult Cells Dendritic Cells Affect Onset and Progress of Psoriasis Diabetes Researchers Find Faster Way to Create Insulin-Producing Cells Human Nasal Epithelial Cells Cultured on a Microchip React to Air Pollutants Why Age Reduces Our Stem Cells' Ability to Repair Muscle Salamander Skin Peptide Promotes Quick and Effective Wound Healing in Mice Living Organ in Animal Grown From Lab-Created Cells Transplanted Stem Cells Help Prevent Bladder Fibrosis After Spinal Cord Injury How Nerve Cells Communicate With Each Other Over Long Distances The Future of Regenerative Medicine A Hybrid-Motor Helps Cells Push Their Way Through Tissues From Rectal Cells to Neurons: Keys to Understanding Cell Transdifferentiation Discovery About Wound Healing Key to Understanding Cell Movement Researchers Seek 'Safety Lock' Against Tumor Growth after Stem Cell Transplantation Implanted Neurons Become Part of the Brain Common Variation Genes Behind the Risk of Autism Stem Cells from Nerves Form Teeth Piggy-Backing Cells Hold Clue to Skin Cancer Growth Age-Related Macular Degeneration Occurs Much Earlier Than Previously Assumed Experimental 'Pulse Radiotherapy' Kills Cancer Cells While Sparing Healthy Tissue Opening-Up the Stem Cell Niche Blocking Cells' Movement to Stop the Spread of Cancer Enlightening Cancer Cells Cancer Uses Stem Cells as a Shield to Escape Drug Attacks Spanish Scientists Grow Artificial Skin Using Umbilical Stem Cells Researchers Identify Main Genes Responsible for Asthma Attacks in Children Fatty Acid Produced by Gut Bacteria Boosts the Immune System Carbon Dioxides New-Found Signaling Role Could be Applied to Blood Flow, Birth, and Deafness New Discovery on Early Immune System Development Cow Milk Peptide Kills Stomach Cancer Cells Dental Pulp Stem Cells Promote the Survival and Regeneration of Retinal Cells After Injury Inducing Visual Function How Cells Translate Signals from Surroundings into Internal Signals Major New Role for B Cells Discovered New Genes for Regulating the Heart Rhythm Discovered Nanotechnology Adds Bling to Antibacterials Invisibility Cloak for Immune Cells New Material Improves Wound Healing, Keeps Bacteria from Sticking Pushing Cells Towards a Higher Pluripotency State Stem Cell Transplantation for Severe Sclerosis Associated With Improved Long-Term Survival Microenvironment of Hematopoietic Stem Cells Can Be a Target for Myeloproliferative Disorders Stem Cell Mobilization Therapy May Effectively Treat Osteoarthritis A Roadmap for Pure Organ Cells Stem Cells: The Fat Source Makes the Difference Transfer of a Few Immune Cells Can Protect Immunodeficient Patients Malaria: Blood Cells Behaving Badly Our Own Treacherous Immune Genes Can Cause Cancer after Viral Infection Potential New Treatment May Protect Celiac Patients from Gluten-Induced Injury Alcohol Abuse Damage in Neurons at a Molecular Scale Identified for First Time Immune Cells Found Near Tumors Boost Breast Cancer Survival Cancer Cells Do It the "Quick-and-Dirty Way" Stem Cells Are a Soft Touch for Nano-Engineered Biomaterials Stem Cells Found to Play Restorative Role When Affecting Brain Signaling Process Healthy Tissue Grafted to Brains of Huntington's Patients Also Develops the Disease Unlocking the Potential of Stem Cells to Repair Brain Damage Dying Cells Trigger Immunity Nanotechnology Takes on Diabetes Blood Vessel Research Offers Insights into New Treatments for Eye Diseases Migrating Stem Cells Possible New Focus for Stroke Treatment A New Molecule for High-Resolution Cell Imaging Functional Nerve Cells from Skin Cells Genetic Tracking Identifies Cancer Stem Cells in Human Patients Nanogel Heals Burn Wounds Faster Understanding Aspirin's Effect on Wound Healing Offers Hope for Treating Chronic Wounds How Immune Cells Use Steroids 'Parent' Cells Reset the Cell Division Clock Single Treatment for AMD 'Unlikely' One Step Closer to Cell Reprogramming DNA Nanotechnology Clamping Down on Cancer-Causing Mutations Stem Cell Production Becomes Much Easier With Polymer-Based 3D Culture System Study Offers Evidence Supporting Use of Autologous Stem Cell-Based Treatment for Stroke How Do Our Cells Move? Liquid Droplets Could Explain Stem Cells from Teeth Can Make Brain-Like Cells Loss of Memory in Alzheimer's Mice Models Reversed Through Gene Therapy Bionic Ear Technology Used For Gene Therapy Scientists Discover a New Way to Enhance Nerve Growth Following Injury 'Dustman' Protein Helps Bin Cancer Cells Researchers Discover New Marker Derived From Human Umbilical Cord Blood Researchers Transplant Regenerated Esophagus Glaucoma Can Actually Be the Diabetes of the Brain, Say Researchers Virus-Fighting Genes Linked to Mutations in Cancer 3-D Printing Cancer Cells to Mimic Tumors Scientists Grow Cartilage to Reconstruct Nose Two New Genes Linked to Intellectual Disability Nanotechnology 'Smart Band-Aid' Monitors Movement of Disorder Patients The Multiplication of Cells under Close Observation p53 Cuts Off Invading Cancer Cells GDNF Transfection Promotes Neuronal Differentiation of Bone Marrow Mesenchymal Stem Cells New Approach Makes Cancer Cells Explode Scientists Create Stem Cells from a Drop of Blood from a Finger Prick Inflammation Mobilizes Tumor Cells Stem Cell Gene Therapy: Chances and Risks New Stem Cell Research Removes Reliance on Human and Animal Cells Could Sticky Cells Cause Drug Resistance in Chronic Blood Cancer? B-Cells Aggravate Autoimmune Diseases Exploring How Cancerous and Healthy Cells Compete May Help Us Understand Cancer Photopharmacology: Optoswitches Turn Pain Off and Sight On Dual Role of Brain Glycogen Revealed by Researchers Dangerous Mistaken Identity: Protein Folding Leads to Some Neurodegenerative Diseases Gene Therapy for Parkinson's Disease Yields Promising Results in First Patients Lab Grown Cells Offer Hope for New Liver Disease Treatments Research into Fruit Fly Cells Could Lead to Cancer Insights Gene Therapy for Human Skin Disease Produces Long-Term Benefits Stem Cell Replacement for Common Age-Related Blindness Patient's Own Bone-Marrow Stem Cells Could Treat Resistant TB Genetic Screening Identifies Genes Driving Resistance with a Guide RNA Library Transitioning Epithelial Cells to Mesenchymal Cells Enhances Cardiac Protectivity Combination of Cell Transplantation and Gene Therapy for Alzheimer's Disease Researchers Find Potential New Treatment Approach For Pancreatic Cancer Young Killer Cells Protect Against Infectious Mononucleosis 3-D Tissue Printing: Cells from the Eye Inkjet-Printed for the First Time Liver Cells Benefit From Mesenchymal Stem Cell Co-Culture Prior to Transplantation New Hope for Stem Cells, Regenerative Medicine Emerges From the Lab Will Stem Cell Therapy Help Cure Spinal Cord Injury? Brittle-Bone Babies Helped by Fetal Stem Cell Grafts Skin Cells Can Migrate Over Regions Devoid of Support From Extracellular Matrix Skin's Own Cells Offer Hope for New Ways to Repair Wounds, Reduce Impact of Aging on the Skin How the Cells Remove Copper Precise Docking Sites for Cells Antivirals for HCV Improve Kidney, Cardiovascular Diseases in Diabetic Patients Researchers Uncover Mechanism Controlling Tourette Syndrome Tics Norfolk Islanders' Genes Yield Bounty of Insight Into Heart Disease: Study Human Stem Cells Predict Efficacy of Alzheimer Drugs Malignant and Healthy Cells Display Characteristic Fractal Patterns Simulating New Treatment for Retinal Degeneration The Heart's Own Stem Cells Play Their Part in Regeneration Bone Grafting Improvements with the Help of Sea Coral Attractants Prevent Nerve Cell Migration What Composes the Human Heart? Protein Coding Junk Genes May Be Linked to Cancer "Kissing Genes" Breakthrough Are Probiotics a Promising Treatment Strategy for Depression? Redesigned Protein Opens Door for Safer Gene Therapy Healing Powers of Cells Scientists Decipher How the Immune System Induces Liver Damage During Hepatitis Bacterial Toxin Sets the Course for Infection Important Step Towards Stem Cell-Based Treatment For Stroke Unlocking a Secret of Stem Cell Stability Researchers Expose Molecular Secrets of Bile Duct Cancers From Different Countries How Nanotechnology Can Advance Regenerative Medicine Predicting the Fate of Stem Cells Antibody Fragment Effectively Fights Alzheimer's Disease in Mice New Treatment Potential for Severe Asthma Patients Nanopore Made for Gene Therapy, Drug Delivery Stem Cell Transplant Repairs Damaged Gut of Inflammatory Bowel Disease Stem Cells Converted to Living Intestinal Patches Rapid Blood Test to Diagnose Sepsis at the Bedside Could Save Thousands of Lives New Way to Treat Chronic Kidney Disease and Heart Failure Cancer Cells' Communication Path Blocked A Holistic Approach Catches Eye Disease Early How a Ubiquitous Herpes Virus Sometimes Leads to Cancer Mutated Stem Cells Trigger Pituitary Tumors in Children Study Explains Why Diabetic Retinopathy Is Difficult to Treat Battling Defiant Leukemia Cells Sieving Through 'Junk' DNA Reveals Disease-Causing Genetic Mutations Specialized Intestinal Cells Cause Some Cases of Crohn's Disease Leukemia Cells Are Addicted to a Healthy Gene Patient's Own Cells Might Be Used as Treatment for Parkinson's Disease Without a Trace: Cells Keep to One Direction by Erasing the Path 'X-Shape' Not True Picture of Chromosome Structure, New Imaging Technique Reveals With Carbon Nanotubes, a Path to Flexible, Low-Cost Sensors: Potential Applications Range from Air-Quality Monitors to Electronic Skin Early Test Warns of World's Leading Eye Disease A New Method for Analyzing Gene Expression in Single Cells Opens a Window into Tumors and Other Tissues Immune Cells Open Window to Breast Cancer Risk Scientists Produce Induced Pluripotent Stem Cells by Removing One Protein Pancreatic Stem Cells Isolated From Mice Specific Sugar Molecule Causes Growth of Cancer Cells Fate of New Genes Cannot Be Predicted Embryonic Stem Cells Produced in Living Adult Organisms Researchers Investigate Importance of STAT1 Phosphorylation in NK Cells Automated Method Could Prevent Blindness by Detecting Glaucoma in Its Early Stages Singapore Scientists Discover New RNA Processing Pathway Important in Human Embryonic Stem Cells Moving Genes Have Scientists Seeing Spots Clingy Platelets Suggest Potential Treatment Strategy for Rheumatoid Arthritis How Brain Microcircuits Integrate Information from Different Senses 'Safe' Levels of Environmental Pollution May Have Long-Term Health Consequences Newly Discovered Weakness in Cancer Cells Make Them More Susceptible to Chemotherapy Intestinal Flora Determines Health of Obese People Cancer Scientists Discover Novel Way Gene Controls Stem Cell Self-Renewal Migraine May Permanently Change Brain Structure BES Procedure May Prevent Cell Apoptosis in Stem Cell-Based Transplantation Therapy Listening to Blood Cells: Simple Test Could Use Sound Waves for Diagnosing Blood-Related Diseases 'Mini Human Brains' Created: Scientists Grow Human Brain Tissue in 3-D Culture System Immunology: White Blood Cells Show Their Stripes Omega-3 Reduces ADHD Symptoms in Rats Nanotechnology Safety: New Tests for Determining Health and Environmental Effects An Organized Approach to 3D Tissue Engineering Discovery of Novel Gene Mutations in Leukemia Patients Opens Up Personalized Therapy Options Impaired Autophagy Associated With Age-Related Macular Degeneration Genetic Associations for Gastrointestinal Condition in Infants Cancer's Origins Revealed Heart's Own Stem Cells Offer Hope For New Treatment of Heart Failure Shining Stem Cells Reveals How Our Skin is Maintained New Biomarker Could Reveal Alzheimer's Disease Years Before Onset New Type of Blood Stem Cell Could Help Solve Platelet Shortage New Gene Therapy Advance Holds Promise for the Treatment of the Global Killer HBV New Research Suggests Glaucoma Screenings for Sleep Apnea Sufferers High-Tech Laser Technique Offers Control of Gene Therapy and Genetic Engineering Method Rheumatoid Arthritis Heightens Risk of Dangerous Leg and Lung Blood Clots High Pain Sensitivity Linked to Dry Eye Disease Symptoms Boning Up: Researchers Find Best Stem Cells for Bone Marrow Transplants How to Achieve a Well Balanced Gut: 'Peacekeeper' in the Gut Identified Breast Cancer Cells' Sugar Craving Is Target for New Type of Treatment Eucalypts Give Nano-Medicine a Golden Boost Could Sleeping Stem Cells Hold Key to Treatment of Aggressive Blood Cancer? New Drugs to Find the Right Target to Fight Alzheimer's Disease Bio-Imaging: Probing For Deeper Diagnostics Scientists Successfully Generate 'Artificial Bones' From Umbilical Cord Stem Cells Key Target Responsible for Triggering Detrimental Effects in Brain Trauma Identified New Stem Cell Gene Therapy Gives Hope to Prevent Inherited Neurological Disease Heading for Regeneration: Researchers Reactivate Head Regeneration in Regeneration-Deficient Species of Planarians Between B Cells and T Cells When Cells Are Consumed by Wanderlust Stem Cell Advance Boosts Prospects for Retina Treatment Protein Responsible For 'Bad' Blood Vessel Growth Discovered New Techniques Use Lasers, LEDs, and Optics to 'See' Under the Skin A Step Forward in Neuronal Regeneration Movement Without Muscles Study in Insects Could Inspire Robot and Prosthetic Limb Developments Stem Cell Research Reveals Clues to Brain Disease Gene Transfer KOs Two Inherited Illnesses Unique Route to Stem Cells: Designer Proteins Developed to Deliver Stem Cells Gene Therapy Using Lentivirus Promising in Three Youngsters Improved Outlook For Immune-Based Therapies Cells Make Costume Changes for Cardiac Regeneration New Tissue Engineering Breakthrough Encourages Nerve Repair How Quality Control Works in Our Cells Bacteria Communicate to Help Each Other Resist Antibiotics Proteins Key in Stem Cell Production Identified Technological Breakthrough Paves the Way for Better Drugs The Role of Noncoding 5S rRNA in Protecting the P53 Tumor Suppressor Gene Discovered Nanotechnology Helps to Test Antibiotic Treatment in Minutes Gene Therapy Cures a Severe Pediatric Neurodegenerative Disease in Animal Models Diabetic Retinopathy Severity Inversely Tied to Cognition Olfactory Ensheathing Cells Treatment Improves Neurological Function in SCI Patients: Study Transplanted Bone Marrow Stem Cells in Patients with Type 2 Diabetes Reduce Insulin Use Silver Could Promote Colonization of Bacteria on Medical Devices Scientists One Step Closer to Preventing Bacterial Infections of Implanted Medical Devices No Danger of Cancer Through Gene Therapy Virus Nanotechnology Helps Track and Improve Drug Action in Pancreatic Cancer Enzyme Could Be the Key to Aiding Wound Healing in Diabetic and Elderly People Researchers Identify Ways to Inhibit Function of Key Protein Linked to Stem Cell-Like Behavior in CML CSIRO Develops Test to Improve Stem Cell Safety Rabbit Wears Contact Lenses with Light-Emitting Diode: New Class of Transparent, Stretchable Electrodes New Possibilities for Prostate Cancer Treatment New Weapon in Fight Against Cervical Cancer Trichloroethylene Exposure Linked to Increased Risk of Some Cancers Gene Therapies for Regenerative Surgery Are Getting Closer Identifying Abnormally Activated Genes to Predict Cancer Virulence First Successful Treatment of Pediatric Cerebral Palsy with Autologous Cord Blood Stem Cells from Fat Outperform Those from Bone Marrow in Fighting Disease Scientists Make Breast Cancer Advance That Turns Previous Thinking on Its Head Mapping a Route to Stem Cell Therapies New Study Pinpoints Biochemical Mechanism Underlying Fibrosis Following Glaucoma Surgery Early Screenings Key to Diagnosing Glaucoma Blood-Producing Stem Cells Created From iPS Cells Accelerated Aging in Children: Promising Treatment for Progeria Within Reach Researchers Reveal How Genes Are Turned On and Off During Early Human Development Scientists Make Key Stem Cell Discovery Schizophrenia May Be Diagnosed by Testing Nerve Cells from the Nose Clarifying the Effect of Stem Cell Therapy on Cancer Cell Infusion No Help After STEMI Tissue Engineering Combined With Gene Therapy Can Manage Repair of Defective Articular Cartilage Examining Function of All Genes in Zebrafish Genome Could Benefit Human Health, Researchers Say Metastasis Stem Cells in the Blood of Breast Cancer Patients Discovered MicroRNA HCV Therapy Has Big Promise Stem Cells Enable Personalized Treatment for Bleeding Disorder 3D Printer Can Build Synthetic Tissues Scientists Form New Nerve Cells -- Directly in the Brain Smelling Genetic Information: Molecules Allow Mice to Sniff Out the Genes of Other Mice Old Mice, Young Blood: Rejuvenating Blood of Mice by Reprogramming Stem Cells That Produce Blood Insights Into the Immune System, From the Fates of Individual T Cells Amniotic Fluid Stem Cells Repair Gut Damage Harnessing Immune Cells' Adaptability to Design an Effective HIV Vaccine Key Step in Manufacture of Red Blood Cells Decoded: Subtle Regulatory Chords Direct the Birth of Blood Cells Third-Party Blood Stem Cell Transplantation as a Factor to Impact on Poor Graft Function Fasting Time for Tumor Cells Asterixs Roman Foes: Analogy for How Cancer Cells Move and Grow Virus and Genes Involved in Causation of Schizophrenia Epigenetics: Neurons Remember Because They Move Genes in Space Lipid Nanoparticles Are Ideal For Delivering Genes and Drugs, Researchers Show Pre-Clinical Research Shows Promising Treatment for Diabetic Wounds Using Stem Cells Switch Critical to Wound Healing Discovered RNA Interference Identifies Function of Genes Linked With Cardiovascular Disease How to Mend a Broken Heart: Advances in Parthenogenic Stem Cells Sweet News for Stem Cell's 'Holy Grail' Small Groups of Brain Cells Store Concepts for Memory Formation -- From Luke Skywalker to Your Grandmother Reversion of Adult Cells into Pluripotent Stem Cells Occurs via Nanog and its Partners Tet1 and Tet2 Implants Make Light Work of Fixing Broken Bones First Signals from Brain Nerve Cells with Ultrathin Nanowires Retinal Implants with Wireless Microchip Restore Functional Vision in Retinitis Pigmentosa Patients, Research Finds Stem Cell-Based Bioartificial Tissues and Organs New Study Shows Stem Cells Promise as Future ALS Treatment Stem Cell Discovery Gives Insight Into Motor Neuron Disease Genes for Autism and Schizophrenia Only Active in Developing Brains New Genes for Short-Sightedness: 24 New Genes That Cause Refractive Errors and Myopia Identified Reactions in Living Cells Possible to Measure Without Disturbance UAB Researchers Cure Type 1 Diabetes in Dogs Fbxl10 Molecule Plays a Crucial Role in Differentiation of Embryonic Stem Cells Key Protein Revealed as Trigger for Stem Cell Development Breast Cancer Stem Cells Can be Targeted to Stop Disease Neural Stem Cell Transplantation Shows Promise Against Gastrointestinal Motility Disorders A Step Towards Repairing the Central Nervous System Programming Cells: Importance of the Envelope Glial Cells Assist in the Repair of Injured Nerves Stem Cells Aid Recovery from Stroke, Study Suggests Helping Healthy Cells Could be Key to Fighting Leukemia, Research Suggests Tuning Stem Cell Fate: Researchers Uncover Epigenetic Mechanisms of Embryonic Stem Cell Pluripotency and Differentiation Regular Aspirin Use Linked to Increased Risk of Age-Related Macular Degeneration 'Quadruple Helix' DNA Discovered in Human Cells Bacteria's Hidden Skill Could Pave Way for Stem Cell Treatments Sniffing Immune Cells: Immune Cells on the Move Are Guided by Touch and Smell Stem Cell Materials Could Boost Research into Key Diseases Mature T Cells Can Switch Function to Better Tackle Infection Singapore Scientists Discover Genes Responsible for Cornea Blindness Not All Stem Cells Are Equally Efficient For Use in Regenerative Medicine Living Cells Behave Like Fluid-Filled Sponges Human Genes Influence Gut Microbial Composition, Study Suggests New Stem Cell Approach for Blindness Successful in Mice Early Predictor for Glaucoma Identified Stem Cell Technology Could Help Harness Patients' Own Immune Cells to Fight Disease Cellular Metabolism Arms T Cells to Battle Viruses and Tumors New Technology Allows Scientists to Capture and Preserve Cancer Cells Circulating in the Bloodstream More Than 200 Genes Identified for Crohn's Disease The End of a Dogma: Bipolar Cells Generate Action Potentials Young Scientist Helps Identify Cause of Widespread Eye Disease Researchers Reveal Role of Dendritic Cells in Type 1 Diabetes Autoimmune Disease: Retraining White Blood Cells Team Studies Role of White Blood Cells in Kidney Failure Study Shows Starving Cancer Cells of Key Nutrient Slows Tumor Growth Cancer Stem Cells Isolated From Kidney Tumors Delivering Stem Cells by Post Means More Effective Global Treatment New SARS-Like Virus Infects Both Human and Animal Cells Salmonella Spreads by Targeting Cells in Our Gut, Study Shows Valuable Tool for Predicting Pain Genes in People: 'Network Map' of Genes Involved in Pain Perception Cocktail Boosts Immune Cells in Fighting Cancer A Relationship Between Cancer Genes and the Reprogramming Gene SOX2 Discovered Rapid Eye Movements Significantly Delayed in People with Glaucoma Inducing Stem Cells to Become Different Cell Types Efficiently Now Possible Using a Three-Dimensional Platform Study Reports Breakthrough in the Understanding of Embryonic Stem Cells How Different Nerve Cells Develop in the Eye: Development in Fish Embryos with Aid of 4D Recordings Stem Cell-Derived Dopaminergic Neurons Rescue Motor Defects in Parkinsonian Monkeys Stopping the Flow of Cancer Cells Promotes the Spread of Tumors Communication Channel Between Cells and Machines Paves Way Toward Bio-Hybrid Robots Fountain of Youth Technique Rejuvenates Aging Stem Cells Brain Cells Made from Urine Cocktail Boosts Immune Cells in Fighting Cancer Stem Cells to Aid Search for New Drugs in Hard-to-Treat Conditions Stem Cell-Derived Dopaminergic Neurons Rescue Motor Defects in Parkinsonian Monkeys Topical Simvastatin Shown to Accelerate Wound Healing in Diabetes Study Suggests Different Organ-Derived Stem Cell Injections Improve Heart Function Scientists Discover New Method of Gene Identification Newly Discovered Effects of Vitamin D on Cancer Stem Cells Develop Best in 3D A Novel Therapeutic Advancement in the Search for Heart Muscle Progenitor Cells Hope for Heart Attack Patients First Randomized Controlled Trial to Show Spinal Cord Regeneration in Dogs Stem Cell Clinical Trials to Tackle Diabetes Bioprinting Has Promising Future Nanokey Opens Tumors to Attack Stem Cell Scientists Discover Potential Way to Expand Cells for Use With Patients A Secret of Tumor Stem Cell Survival: Scientists Make Progress Against a Devastating Cancer Scientists Identify Progenitor Cells, Potential New 'Roots' of Breast Cancer How to Make Stem Cells Nuclear Reprogramming Moves a Step Forward Researchers Discover Turbo Switch of Calcium Pump in Biological Cells Watching the Cogwheels of the Biological Clock in Living Cells Breast Cancer Cells Spread by Digging Their Escape Route Finnish Researchers' Discover New Blood-Vessel-Generating Cell with Therapeutic Potential Tissue-Engineered Kidneys: Researchers Make Important Strides Hormone-Producing Thyroid Grown From Embryonic Stem Cells Zinc Fingers: A New Tool to Fight Against Huntington's Disease Brain Tumors: Artificial Stimulation of the Immune System Could Mean Less Aggressive Treatment Researchers Discover How Melanoma Cells Circumvent the Immune System Two Genes Linked to Increased Risk of Diabetes-Associated Kidney Disease Sperm and Eggs Created in Dish Produce Mouse Pups New Human Neurons from Adult Cells Right There in the Brain Adult Stem Cells Change Their Epigenome to Generate New Organs Mouse Pancreatic Stem Cells Successfully Differentiate Into Insulin Producing Cells New Gene-Therapy Approach Could Improve Obesity Treatment Cloned Receptor Paves Way for New Breast and Prostate Cancer Treatment Cloned Receptor Paves Way for New Breast and Prostate Cancer Treatment Human Embryonic Stem Cells Could Help to Treat Deafness Genetic Test Predicts Risk for Autism Spectrum Disorder Newly Identified Stem Cell Population in Skin's Epidermis Responsible For Tissue Repair Protecting Genes, One Molecule at a Time Singapore Scientists Find Genes Associated With Glaucoma SickKids First to Grow Lung Cells Using Stem Cell Technology New Insights into Salt Transport in the Kidney Green Tea Compound Shows Promise for Tackling Cancer New Pre-clinical Results: Ex Vivo Applications of RNAi to Enhance Dendritic Cell Vaccines for Cancer Researchers Identify Which Sensory Nerve Cells Contribute to Chronic Nerve Pain Stem Cell Therapy Shows Promise in Repairing Stress Urinary Incontinence Viruses With Integrated Gene Switch Genes Carried by E. Coli Bacteria Linked to Colon Cancer Stem Cells Thrive on Superficial Relationships Nanomaterials Help the Heart to Heal Blood Cancer Cells Initiate Signaling Cascade From Microns to Centimeters: Researchers Invent New Tissue Engineering Tool Breakthrough Treatment Reduces Post-Surgical Scarring For Glaucoma Patients Vitamin-Boosted Stem Cells Show Promise in Curing Baldness Dr. Peter Wearden Provides International Assistance with Regenerative Medicine Medical Device New Gene Therapy Strategy Boosts Levels of Deficient Protein in Friedreichs Ataxia Regenerating Bone Tissue Using Gene Therapy: New Method May Work With Other Tissues as Well Microwave Heating Improves Artificial Bone Mechanisms that Allow Embryonic Stem Cells to Become Any Cell in the Human Body Identified Single-Cell Parasites Co-Opt 'Ready-Made' Genes from Host: Study Stem Cell Research Aids Understanding of Cancer Finished Heart Switches Stem Cells Off New Target Discovered Deep Within Cancer Cells How Tumor Cells Create Their Own Pathways An Economical, Effective, and Biocompatible Gene Therapy Strategy Promotes Cardiac Repair Origins of Key Immune Cells Discovered Amniotic Fluid Yields Alternatives to Embryonic Stem Cells New Study Finds Genetic Causes of Osteoarthritis Researchers Prevent Mice from Developing Diabetes Diabetes Drug Makes Brain Cells Grow Apolipoproteins Are Biomarkers for Diabetic Retinopathy Diabetes Reversed in Mice Using Stem Cells Magnet Helps Target Transplanted Iron-Loaded Cells to Key Areas of Heart Stem Cell Transplantation into Mouse Cochlea May Impact Future Hearing Loss Therapies Anchoring Points Determine Fate of Stem Cells European Researchers Crack Embryonic Stem Cells Mystery Self-Organized Retinal Tissue from Human Embryonic Stem Cells Regenerated Cells May Restore Vision After Corneal Dysfunction Japanese Scientists Show 'New' Liver Generation Using Hepatocyte Cell Transplantation Export Extravaganza in Human Cells Anti-Psychotic Drug Pushes Cancer Stem Cells Over the Edge Stem Cell Researchers Map New Knowledge About Insulin Production Stem Cell Research Paves Way for Progress on Dealing With Fragile X Retardation Nine New Breast Cancer Risk Genes: Landscape of Cancer Genes and Mutational Processes in Breast Cancer Complicated Prosthetic Retina Offers Simple Solution for Restoring Sight Patients' Skin Cells Turned Into Heart Muscle Cells to Repair Their Damaged Hearts Human Neural Stem Cells with Tumor Targeting Ability Discovered New Stem Cell Found in Brain: Finding Could Be Key to Developing Methods to Heal and Repair Brain Injury and Disease Photoreceptor Transplant Restores Vision in Mice Hair Regeneration from Adult Stem Cells Key to New Antibiotics Could Be Deep Within Isolated Cave Skin and Umbilical Cord Cells Turned Directly Into Nerve Cells Growing Where No Cell Has Grown Before Gene Gives Cells a 'Safety Belt' Against Genetic Damage Trinity Researchers Report Major Eye Disease Breakthrough Asthma: A Vaccination That Works Using Intramuscular Injection Defect in Transport System Causes DNA Chaos in Red Blood Cells Scientists Shed Light on How Liver Repairs Itself REST Molecule Acts as an Adapter in Stem Cells New Zealand Scientists Discover How Infection Triggers Blood Stem Cell Growth Engineering Bowel Transplants Somatic Stem Cells Obtained From Skin Cells for First Time Ever Double Whammy: RNAi Enhances Lung Cancer Therapy Largest Gene Therapy Trial for Cystic Fibrosis Begins New Lab Model of Alzheimers Disease Will Help Research A New Short Cut for Stem Cell Programming Stem Cell Study Aids Quest for Motor Neuron Disease Therapies Neurons Change Shape After Gene Therapy Research Breakthrough Shows Promise in Offsetting Cornea Transplant Rejection Researchers Discover a Switch That Controls Stem Cell Pluripotency UBC-Vancouver Coastal Health Researcher Discovers New Type of Spinal Cord Stem Cell Rare Immune Cells Could Hold Key to Treating Immune Disorders Heart Failure's Effects in Cells Can Be Reversed with a Rest Osteoporosis Drugs May Increase Risk of Serious Eye Disease Artificial Thymus Tissue Enables Maturation of Immune Cells Cheek Progenitor Cells Found to Be Vastly More Potent at Immunosuppression than Bone Marrow MSCs How Cells Brace Themselves for Starvation Live From the Thymus: T-Cells On the Move Bioengineering to Repair and Generate Healthy Skin A Recipe for Isolating and Growing Human Intestinal Stem Cells at the Lab Bench Synthetic Protein Amplifies Genes Needed for Stem Cells Few Genes Control Neuronal Function Human Stem Cells Improve Vision in Rats With Glaucoma-Like Nerve Cell Damage Scientists Make Groundbreaking Discovery on Stem Cell Regulation British Study May Improve Glaucoma Assessment and Treatment Clean Correction of a Patient's Genetic Mutation High Level of Stem Cells in the Blood May Be Associated With Favorable Prognosis for Myeloma Patients Stem Cell Research Could Prevent Premature Births Dental Pulp Stem Cells Transformed by 'Bad Breath Chemical Stem Cell Approach Primes Immune System to Fight Cancer Reprogramming Stem Cells to a More Basic Form Results in More Effective Transplant, Study Shows Erasing the Signs of Aging in Cells is Now a Reality UQ Researchers Make Breakthrough in Stem Cell Research Researchers Discover Key to Immune Cell's 'Internal Guidance' System Queensland Researchers Make Cancer Treatment Breakthrough Stem Cells Can Repair a Damaged Cornea Report Shows Risk of Blindness Halved Over Last Decade Lithocholic Acid Kills Cancer Cells But Not Healthy Cells Collaborative Research Sheds Light on New Cancer Stem Cell Therapies Therapeutically Useful Stem Cell Derivatives in Need of Stability Scientists Shed New Light on Link Between 'Killer Cells' and Diabetes Naturally Produced Protein Could Boost Brain Repair Hopes for Reversing Age-Associated Effects in MS Patients Aging-Related Degeneration Can Be Caused by Defects of Energy Metabolism in Tissue Stem Cells Three New Eczema Genes Discovered Bone Marrow-Derived Cells Differentiate in the Brain Through Mechanisms of Plasticity No Brakes on Breast Cancer Cells Gene Therapy Achieves Early Success Against Hereditary Bleeding Disorder Scalable Amounts of Liver and Pancreas Precursor Cells Created Using New Stem Cell Production Method New Discovery of Adult Stem Cells Gives Fresh Possibilities For Heart Regeneration Genome-Wide Study into New Gene Functions in the Formation of Platelets Stabilizing Entangled Spaghetti-Like Materials: Controlling Forces Between Oppositely Charged Polymers Opens New Route For Gene Therapy Vectors Singapore Scientists Lead Human Embryonic Stem Cell Study to Advance the Field of Regenerative Medicine Research Researchers Decode a Puzzling Movement Disorder Stem-Cell-Seeded Bioartificial Tracheal Transplant Feasible New Muscle Repair Gene Discovered MedUni Vienna: Key Gene Function Against Cell Death Discovered Jamb and Jamc Are Essential Proteins for the Fusion of Muscle Cells: Study Alzheimer Brain Destroyed By Too Many Blood Vessels Lasers Light the Path to Neuron Regeneration Scientists Announce Human Intestinal Stem Cell 'Breakthrough' For Regenerative Medicine Scientists Make Historic Breakthrough in Cancer Therapy Trial Scientists Discover How Cancers Generate Muscle-Like Contractions to Spread Around the Body Discovery Could Create Retinas From 'Jell-O' New Research Validates Clinical Importance of Leukemia Stem Cells Mural Cells from Saphenous Vein Could Have Long-Term Benefits in Heart Attacks 'Open Wide' For New Stem Cell Potential: Stem Cells of the Oral Mucosa Stay Young Research Discovers Frequent Mutations of Chromatin Remodeling Genes in TCC of the Bladder New Resource to Unlock the Role of MicroRNAs Chained Muscle Tissue Culture Develops Blood Vessels and Aligned Structure Researchers Combine Stem Cell with Artificial Chromosome to Treat Duchenne Muscular Dystrophy in Mice Insect Virus Holds the Key to Safer Stem Cell Therapy Protein Switch Controls How Stem Cells Turn Into New Heart Tissue Stem Cell Breakthrough Heralds New Era of Therapy Development Stem Cell "Memory" Can Boost Insulin Levels Genetic Study Sheds New Light on Ankylosing Spondylitis "Genetic Dimmer Switch" Regulates Wiring in the Developing Brain Gene Therapy Stimulates Protein That Blocks Immune Attack and Prevents Type 1 Diabetes in Mice The Forces of Attraction: How Cells Change Direction UBC Researchers Invent New Drug Delivery Device to Treat Diabetes-Related Vision Loss Serum-Free Cultures Help Transplanted MSCs Improve Efficacy Singapore Scientists Discover How to Control Fate of Stem Cells Blue Light Enables Genes to Turn On Molecular Glue Sticks It to Cancer Nanotubes to the Rescue: Brain Cells Rescued by Gene Silencing Offers New Options for Stroke New Approach to Cancer Vaccines Proves Successful in Early Studies New Genetic Technique Converts Skin Cells Into Brain Cells Heart Has Built-in Repair Mechanism Cancer-Seeking 'Smart Bombs' Target Kidney Cancer Cells Silencing a Deadly Conversation in Breast Cancer MDC Researchers Discover Key Molecule for Stem Cell Pluripotency Scientists Find New Drug Target in Breast Cancer Scientists Boost Antitumor Effects of Dendritic Cell Therapy Using Inflammatory Virus Stem Cell Treatment to Prevent Leukemia Returning is a Step Closer New Drugs Target Delay of Huntington's Symptoms Rare Tumors 'Fingerprint' Used to Develop Cheap and Reliable New Test Cell Therapy Aims to Prevent Transplant Rejection Combo Method Reveals Cells' Signal Systems Team Develops Platform to Monitor Hematopoietic Stem Cells Cell Therapy to Prevent Organ Rejection New Marker Offers Hope for More Reliable Detection of Prostate Cancer Liver Cells Made in the Lab Using Skin Cells: Study Cancer Scientists Discover New Way Breast Cancer Cells Adapt to Environmental Stress The Sweet Mysteries of the Nervous System Discovery Demonstrates Potential MS Therapy Could Kill Brain Cells Researchers Discover Protein That Could Help Prevent the Spread of Cancer Tekmira and the National Cancer Institute Publish Promising Data Demonstrating the Anti-Tumor Activity of a Novel Cancer Target Simple Injection Could Limit Damage from Heart Attacks and Stroke First Successful Transplantation of a Synthetic Tissue-Engineered Windpipe Gene Find Could Aid Nerve Repair Successful Strategy Developed to Regenerate Blood Vessels Tiny Antibody Fragments Raised in Camels Find Drug Targets in Human Breast Cancer Cells Scientists Make Skin Repair Discovery Scientists Identify Key Trigger for Pancreatic Beta Cell Regeneration Researchers Mimic Body's Own Healing Potential to Create Personalized Therapies for Inflammation Womb Cancer Genome Scan Reveals Prostate Cancer Link University of Granada Researchers Make the First Bioartificial Organ in Spain The Self-Made Eye: Formation of Optic Cup from ES Cells Newly Discovered Epigenetic Tag Offers Insight into Embryonic Stem Cell Regulation Periodontal Stem Cell Transplantation Shows Promise Molecular Muscle: Small Parts of a Big Protein Play Key Roles in Building Tissues New Study Pinpoints Why Some Microbial Genes are More Promiscuous Than Others Trapping Prostate Cancer Cells to Keep Them from Spreading Provides Hope Two-Pronged Approach Brings Hope for Bowel Cancer Treatment New Cell Therapy a Promising Atherosclerosis Treatment Gene-Fuelled Transporter Causes Breast Cancer Cells to Self-Destruct Gene Responsible for Severe Osteoporosis Disorder Discovered Genetic Abnormalities Associated With the Creation of Stem Cells Discovered Stem Cell Study Could Aid Motor Neuron Disease Research Scientists Discover Cause of Rare Skin Cancer That Heals Itself Human Umbilical Cord Blood Cells Aid Diabetic Wound Healing Cancer-Related Pathways Reveal Potential Treatment Target for Congenital Heart Disease Personalized Medicine Comes Within Reach Worlds First Chemical Guided Missile Could Be the Answer to Wiping Out Cancer Canadian Researchers First Worldwide to Generate Pluripotent Stem Cells From Horses U of Alberta Researcher Discovers Potential Cancer Therapy Target How Nasal Stem Cells Might Prevent Childhood Deafness Boosting Bodys Immune Response May Hold Key to HIV Cure McMaster Discovery Provides New Insights into Pluripotent Stem Cell Biology Early Infusion of Donor T Regulatory Cells Prevents Graft-Versus-Host Disease and Enhances Immune Recovery in High-Risk Blood Cancer Patients Erg Gene Key to Blood Stem Cell 'Self-Renewal' Reprogrammed Stem Cells Hit a Roadblock Partnership of Genes Affects the Brain's Development Immune Interaction Optimizes Fetal Nourishment During Pregnancy New Anti-HIV Gene Therapy Makes T-Cells Resistant to HIV Infection DMP1 Protein Inhibits Angiogenesis, Could Lead to New Treatments Against Cancer and Other Diseases Research Shows How Pathogenic Bacteria Hide Inside Host Cells Dynamic Systems in Living Cells Break the Rules Uncovering the Trail Behind Growing Too Old, Too Soon Blocking Rogue Gene Could Stop the Spread of Most Cancers Red Blood Cell Hormone Modulates the Immune System Patient-Derived Stem Cells Could Help Test Cardiac Disease Treatments Skin Provides Australia's First Adult Stem Cells for Friedreich Ataxia Nanodisk Gene Therapy Scientists Now Know Why Some Cancers Become Malignant and Others Don't Scientists Bring New Hopes to Translate Human Stem Cell Technologies to the Clinic Scientists Find the Master Switch for Key Immune Cells in Inflammatory Diseases Researchers Find Critical Link Between Down Syndrome and Alzheimers Disease Punctuated Evolution in Cancer Genomes Australian Researchers Identify Gene Mutations That Sabotage Kidney Function Two Genes Linked to Common Gynecological Disease Study Reveals New Possibility of Reversing Damage Caused by MS Some Brain Tumors Mimic the Genetic Program of Germline Cells Leukemia Stem Cell Discovery Degradable Scaffolds for Tissues A Double Block of Blood Vessels to Starve Cancerous Tumors Medicine: Alzheimer's and Heart Attacks Share the Same Genes TB Bacteria Use the Body's Stem Cells to Protect Themselves Study Shows That Blood Stem Cells are Influenced by Their Offspring Fitting a Biological Nanopore into a Man-Made One, New Ways to Analyze DNA New Microscope Reveals Ultrastructure of Cells Transcription Factor Scan Identifies Genetic Cause for Inherited Blindness Overactive FTO Gene Does Cause Overeating and Obesity Team Finds Mutation Involved in Macular Degeneration Origin of Cells Associated With Nerve Repair Discovered Umbilical Cord Cells May Treat Arthritis Rogue Gene Hijacks Stem Cells to Jumpstart Human Cancer Breakthrough in Cancer Vaccine Research ISMETT Celebrates Transplant Number 1000 Pediatric Living-Related Liver Transplant at ISMETT Scientists Find Non-Viral Pathway of Blood Cancer Development How Do We Kill Rogue Cells? Assassin's Tricks Revealed Not All Stem Cells the Same, Say McMaster Researchers Scientists Turn Stem Cells into Cells for Cartilage Repair Scientists Uncover Strategy to Effectively Target Drug Resistant Childhood Cancers Stem Cells Shape Up to Their Surroundings Scientists Reveal Gut Stem Cell Secrets Visionary Stem Cell Technique Offers New Potential to Treat Blindness Scientists Unravel Genetic Risk of Bowel Cancer Scientists 'Clone' Human Virus Responsible for Congenital Malformations and Other Life-Threatening Diseases Major Genetic Risk Factor for the Most Common Form of Glaucoma Discovered Uncovered: New Ovarian Cancer Gene (A Connection Between Cancer and Endometriosis) Twins Help Researchers to Find Genes for Blindness Identifying Stem Cell Suicide Triggers with ROCK Accelerating the Understanding of Personalized Medicine B Vitamins Slow Brain Atrophy in People with Memory Problems Smoking Damages Fertility Scientific Breakthrough to Pave the Way for Human Stem Cell Factories First Genetic Link to Common Migraine Exposed Insulin Resistance, Type 2 Diabetes Linked to Plaques Associated With Alzheimer's Disease Scientists Describe How Physical Environment Influences Stem Cell Development Important Breakthrough in Origins of Aggressive Breast Cancer Toolbox of MiniPromoters for Gene Research and Therapy Unveiled All Genes in One Go Seeing the World With New Eyes: Biosynthetic Corneas Restore Vision in Humans Gene Therapy to Treat Epilepsy a Step Closer Liver Cells Created From Patients' Skin Cells New Stem Cell Discovery a Preliminary Step for Regenerative Medicine Slippery DNA Holds Clues to Fighting Diseases That Progress with Age Otago Researcher Helps Identify Gene Linked to TB Susceptibility in Africans Stem Cells from the Thymus Reprogrammed to Become Skin Stem Cells Starvation Gene Could Make Resistance Futile for Ovarian Cancer Cells Personalized Genome Sequencing in Cancer TreatmentA Major Breakthrough in Care Genetic Signature in TB Patients Sheds Light on Disease Genetic Differences That Make Some People Susceptible to Meningitis Revealed Gene Affecting Chronic Pain Identified--Opens Way for New Treatment Approach Herpes Virus Treats Head and Neck Cancer Patients Manipulating Cells With a Micro-Suction Cup New Discovery Brings Hope to Treatment of Incurable Blood Cancer Scientists 'Reprogram' Mouse Fat Cells Into Clinically Useful Stem Cells Toxic Trio Identified as the Basis of Celiac Disease Bid to Aid Transplant Cancer Patients Gene Find Offers Breast Cancer Hope Potential Cure Discovered for Degenerative Vision Diseases Leading to Terminal Blindness Cells' Grouping Tactic Points to New Cancer Treatments Finding May Pave Way for New Treatments for Muscle Wasting Diseases New Discovery in Nerve Regrowth Tel Aviv University Develops Method for Tracking Adult Stem Cells as They Regress UK Clinical Trial: Stem Cells to Treat Knee Osteoarthritis Stem Cell Transplantation of Therapy-Resistant Chronic Leukemia Successful Reversal of Fortune for Parkinson's Disease Transplant Treatment Potential Cancer Cause Uncovered Revolutionary Medical Dressing Uses Nanotechnology to Fight Infection Prostate Cancer Study's Dramatic Results Stem Cell Therapy May Provide New Approach to Fight Infection 12 New Genes Linked to Type 2 Diabetes Microbial Protein Restores Vision in Blind Animals Identification of a Novel Tumor Suppressor Bees Help to Beat MRSA Bugs Mechanism Explains Complications Associated With Diabetes Stem Cell Breakthrough Offers New Hope for Lung Disease Patients Scientists Identify Three New Genetic Variants That Increase Testicular Cancer Risk Mechanism Behind Three Cancer-Causing Genes in Acute Lymphoblastic Leukemia Rare Genetic Variations Involving Whole Sections of DNA Implicated in Autism Reprogramming Immune System Cells to Produce Natural Killer Cells for Cancer Obstacles to Stem Cell Therapy Cleared Dental Pulp Cells for Stem Cell Banking How the Wrong Genes Are Repressed Down's Syndrome Chromosome Yields More Cancer-Blocking Genes Ancient Viral Invasion That Shaped the Human Genome Found Sleeping Sickness Study Offers Insight Into Human Cells New Therapy Offers Hope to Spinal Muscular Atrophy Patients Scientists Find Genes Associated With Throat Cancer Transplant Tolerance 'Signature' Identified Progress in Exploring New Avenues for Brain Repair New Findings Complicate Use of Stem Cells Study Sheds Light Into the Nature of Embryonic Stem Cells Harnessing the Power of Stem Cells to Unlock the Secrets of Motor Neuron Disease Stem Cells From Adipose Tissue Turned Into Cardiac Myocytes: Achievement Paves Way for Future Heart Disease Therapies Genetic Link to Risk for Diabetes and Weight Gain: New Study Funnel Vision: New Info About How Cells in the Eye Help Guide Light Into the Retina Researchers Discover New Mechanism in Stem Cell Regulation Researchers Agree on Future of MS Stem Cell Research Researchers Design New Biomaterial That Mimics Muscle Elasticity Junk DNA Drives Cancer Growth Researchers Modify Virus to Hunt Down and Wipe Out Cancer Cells Finding Has Direct Impact on Regenerative Medicine and Assisted Reproduction Link: Ovarian Hormone to Breast Stem Cells Growth Bone-Marrow Stem Cells in MS Show Promise Prenatal Smoking Can Lead to Psychiatric Problems in Children Scientists Create Artificial Human Skin with Biomechanical Properties Using Tissue Engineering Chinese Scientists Discover Marker Indicating the Developmental Potential of Stem Cells Major Breakthrough Offers Hope of Preventing Mitochondrial Diseases Substance in Breast Milk Kills Cancer Cells New Discovery Could Speed Up Liver Repair Hormone Sensitivity of Breast Stem Cells Presents Drug Target Insulin-Producing Cells Can Regenerate in Diabetic Mice Gene Therapy Boosts Recovery From Heart Attack Potential to Produce the Large Quantities of Human Embryonic Stem Cells Required for Transplantation Hormone Replacement in Joint Fluid Has Potential Regenerative Effect Tomato Genes Produce Promising Results Against Brain Tumors Using Stem Cells to Mend Damaged Hips Surgeons Transplant New Trachea Into Child Using His Own Stem Cells to Rebuild Airway How to Fortify the Immunity of HIV Patients Gut Flora Genes Dwarf Human Genome Predicting the Fate of Stem Cells Research Builds on Genetic Link to Autism and Schizophrenia How Nerve Cells Grow Genetic Link Between Mammographic Density and Breast Cancer Study: Stem Cells Sabotage Their Own DNA to Produce New Tissues Bacteria Toxic to Wound-Treating Maggots Stem Cells Rescue Nerve Cells by Direct Contact Novel Theory for Mammalian Stem Cell Regulation How to Shoot the Messenger Targeting Cancer Stem Cells in the Lab Did Bacteria Develop Into More Complex Cells Much Earlier in Evolution Than Thought? Scientists Map Epigenome of Human Stem Cells During Development Regenerative Medicine and Transplantations: How the Immune System Produces T Cells 'Junk DNA' Could Spotlight Breast and Bowel Cancer 'Fountain of Youth' for Stem Cells? Metastasis Formation Revealed in Detail in Real Time Acutely Damaged Livers may be Repaired via Transplanted Hepatocytes Leprosy Susceptibility Genes Identified Gene Identified as Cause of Some Forms of Intellectual Disability Stroke Drug Kills Cancer Cells and Leaves Normal Cells Intact Why Cancer Cells Just Won't Die: Researcher Identifies Protein Which Regulates Cell Suicide It Takes Two to Infect: Structural Biologists Shed Light on Mechanism of Invasion Protein Genetic Analysis Helps Dissect Molecular Basis of Cardiovascular Disease Polyphenols and Polyunsaturated Fatty Acids Boost the Birth of New Neurons, Study Finds Umbilical Cord Blood Stem Cell Transplant May Help Lung, Heart Disorders New Understanding About Mechanism for Cell Death After Stroke Leads to Possible Therapy First Reconstitution of an Epidermis from Human Embryonic Stem Cells Scientists Discover Cells That Control Inflammation in Chronic Disease Cross-Country Runabouts: Immune Cells on the Move Paradoxical Protein Might Prevent Cancer Survival of the Healthiest: Selective Eradication of Malignant Cells Dementia: Rare Brain Disorder is Highly Hereditary Gene Therapy Repairs Injured Human Donor Lungs for the First Time Testicular Tumors May Explain Genetic Mutations in Children Adipose-Derived Stem and Regenerative Cells Show Potential in First-in-Man Treatment of Stress Urinary Incontinence Blood Counts Are Clues to Human Disease Research Points to Potential Chink in Cancer's Armor Sophisticated Genetic Engineering Improves Insulin-Producing Beta Cells Cholesterol Necessary for Brain Development, Study Finds Step Forward for Nanotechnology: Controlled Movement of Molecules Retinal Rescue: Cells Derived From Human Embryonic Stem Cells Reverse Retinal Degeneration Discovery of Regulatory Role of Key Molecule: Step Towards Future Gene Therapy to Control Disease Microchip That Can Detect Type and Severity of Cancer Created Alzheimer's Researcher Demonstrates Specific Immune Response to Vaccine How Stem Cells Make Skin Genes Identified may Help Breast Cancer Diagnosis New Role of Vitamin C in Skin Protection Key Protein Controlling Brain Formation Identified Master Gene that Switches on Disease-Fighting Cells Identified by Scientists Why Don't Brain Tumors Respond to Medication? Silenced Genes as Warning Sign of Blood Cancer Think Zinc: Molecular Sensor Could Reveal Zinc's Role in Diseases Parkinson's Disease: When Cells Run Out of Fuel Switching on the Power of Stem Cells Watching Stem Cells Repair the Human Brain Nano-Magnets Guide Stem Cells to Damaged Tissue Scarring Key to Link Between Obesity and Diabetes Researchers Successfully Reverse Multiple Sclerosis in Animals Viral Mimic Induces Melanoma Cells to Digest Themselves Cancer Cells Are Protected by Our Own Immune System Researchers Learn How Blood Cells 'Talk' Stem Cell 'Daughters' Lead to Breast Cancer Discovery of a Mechanism Controlling the Fate of Hematopoietic Stem Cells Placenta-Derived Stem Cells May Help Sufferers of Lung Diseases Critical Link in Cell Death Pathway Revealed Researchers Discover Genetic Circuit That Regulates Behavior of Stem Cells Asperger Syndrome, Autism, and Empathy: Study Links 27 Genes Stem Cells: Environmental Factors Instruct Lineage Choice of Blood Progenitor Cells Circulating Blood Cells are Important Predictors of Cancer Spread in Children Improving Mouse Heart Function Following Heart Attack New Weapon in the War Against HIV-AIDS: Combined Antiviral and Targeted Chemotherapy Cancer is Second Most Frequent Cause of Death in Individuals with Schizophrenia Synthetic Biology: Gene Therapy Gets Under the Skin Second Chance for Dangerous T-Cells How Cancers Spread to the Brain Health Risks of Nanotechnology: How Nanoparticles Can Cause Lung Damage and How the Damage Can Be Blocked Vaporized Viral Vector Shows Promise in Anti-Cancer Gene Therapy New Way to Enhance Stem Cells to Stimulate Muscle Regeneration Fruit Fly Stem Cells Filmed Live Revolutionary Ultrasonic Nanotechnology May Allow Scientists to See Inside Patients Individual Cells World First: Chinese Scientists Create Pluripotent Stem Cells from Pigs Model for New Generation of Blood Vessels Challenged Non-Wovens as Scaffolds for Artificial Tissue Bone Marrow Stem Cell Co-Transplantation Prevents Embryonic Stem Cell Transplant-Associated Tumors Cancer: Detecting, Targeting, and Disabling Tumor Cells, All in One Step White Blood Cells Can Sprout 'Legs' and Move like Millipedes Bones Made from Human Skin Connective Tissue Potential Preventative Therapy for Type 1 Diabetes Upside-Down World: DNA Protecting Protein Helps Cancer Drug to Kill Cells Stem Cell Focus for IBD Wound Healing Diabetes: MicroRNA Protects Beta Cells Human Stem Cells Promote Healing of Diabetic Ulcers Learning Disabilities in Males: Nine New X Chromosome Genes Linked to Learning Disabilities Stem Cell Therapy Grows New Blood Vessels Scientists Synthesize Gold to Shed Light on Cells' Inner Workings Large Quantity of Stem Cells Produced from Small Number of Blood Stem Cells Common Virus Kills Cancer Stem Cells Researchers Reveal How Immune Cells Can Be Harnessed to Target Melanoma Geologists Research Could Provide Breakthrough for Heart Patients Regulatory Molecule for Tumor Formation or Suppression Identified Dendritic Cells Ensure Immune Tolerance Experts Use Nanotech to Deliver Anti-Cancer Genes Stem-Cell Repair Kit for Stroke Secrets of C. Difficile's Protective Shell Revealed Viscosity Changes in Cancer Cells Studied The Making of an Intestinal Stem Cell Tiny Tool to Control Growing Blood Vessels Opens New Potential in Tumor Research A Stem Cell Bandage for Your Knee An Inexhaustible Source of Neural Cells Study: Cell Injections Accelerate Fracture Healing New Genes Involved in Acute Lymphoblastic Leukemia Play Fundamental Role in Prognosis of the Disease Adult Stem Cells Convert Into Embryonic-like Stem Cells, With Single Factor 'Nonsense' in Our Genes: One in 200 Human Genes Superfluous? Surgical Implants Coated with One of 'Nature's Antibiotics' Could Prevent Infection Advancement in Tissue Engineering Promotes Oral Wound Healing Stem Cells Used to Reverse Paralysis in Animals How Cancer Cells Survive a Chemotherapy Drug Inflammation Worsens Danger Due to Atherosclerosis Scientists Successfully Develop Novel Biomaterial to Help Grow New Blood Vessels New Mechanism Regulates Type I Interferon Production in White Blood Cells Bone Marrow Stem Cells Used to Regenerate Skin Scientists Call Up Stem Cell Troops to Repair the Body Using New Drug Treatments Older Women Who Are More Physically Fit Have Better Cognitive Function Key to Future Medical Breakthroughs is Systems Biology, Say Leading European Scientists Brain Birth Defects Successfully Reversed Through Stem Cell Therapy Breakthrough in Understanding Development of Type 1 Diabetes Heart Regenerates After Infarction -- First Trials with Mice Type 1 Diabetes and Celiac Disease Linked Gene May Make Kids Crave Junk Food Between Dormancy and Self-Renewal: Mouse Model Shows Blood Stem Cells in Action New Targeted Fluorescent-Imaging Compound Allows Researchers to Detect Viable Cancer Cells in Mice Dormant Stem Cells for Emergencies Stem Cell Research Hold Great Promise, but Obstacles Remain, Expert Argues The Light Syringe Portuguese Scientists Discover New Mechanism that Regulates Formation of Blood Vessels Immune Cells Reveal Fancy Footwork New Approach Keeps Tumor Cells from Refueling Pure Insulin-Producing Cells Produced in Mice Neurons Derived from Embryonic Stem Cells Restore Muscle Function After Injury Adult Stem Cell Breakthrough: First Tissue-Engineered Trachea Successfully Transplanted Pfizer Launches Global Regenerative Medicine Research Unit A New Laser Technique Seals and Heals Wounds Jumonjd3: A Key for Unlocking Neuronal Stem Cell Fate Intraspinal Implant of Mesenchymal Stem Cells May Not Heal the Demyelinated Spinal Cord Heartbeat Supplements Power to Run Implanted Medical Devices Umbilical Cord Blood May Help Build New Heart Valves Completely Novel Action of Insulin Unveiled Skin Cancer: Designer Molecule Tackles Malignant Cells by Two Completely Different Routes Stem Cell Therapies for Heart Disease One Step Closer Singapore Develops Cell Therapy Treatment for Cancer New Artificial Heart in Preclinical Testing in Europe Cold Virus Found to Manipulate Genes Scientists Trigger Cancer-like Response from Embryonic Stem Cells New Properties of Skin Stem Cells Breast Stem Cell Fate is Regulated by 'Notch' A Key Mechanism Regulating Neural Stem Cell Development is Uncovered Stem Cells From the Testicle: True Alternative to Embryonic Stem Cells Cornea Uses Stem Cells to Repair Itself Scientists Identify New Bone Protecting Protein Treating Vitiligo With a New Cell Therapy Scientists Identify for the First Time the Genes that Cause Blindness Produced by Corneal Oedema New Site Launched for Families and Scientists to Discuss Stem Cell Research Human Embryonic Stem Cell Secretions Minimized Tissue Injury After Heart Attack Gene Expression Profile Shared by Pluripotent Stem Cells Found New Stem Cell Screening Tool Takes Adult Stem Cell Research to New Level Stem Cell Research Compares Methods of Creating Patient-Specific Stem Cells Scientists Reveal Changes to Embryonic Stem Cells Caused by Down Syndrome New Stem Cell Tools to Aid Drug Development Researchers Replicate Insulin-Producing Cells, Providing New Hope for Diabetics New Discovery About Growth Factor can be Breakthrough for Cancer Research Blood Vessel Cells are Instructed to Form Tube-Like Structures Animal Testing Could Soon Be a Thing of the Past Cells in Eye Could Help Control Sleep Trial: Tumor Regressions Achieved in Relapsed Lymphoma Patients Potential New Treatment for Alzheimers Disease Robot with a Biological Brain: New Research Provides Insights into How the Brain Works SickKids Scientists Uncover the Key to Controlling How Stem Cells Develop Building the Bionic Body Human Embryonic Stem Cells Developed from 4-Cell Embryo--World First May Lessen Ethical Concerns Bath Researchers Study How Cancer Cells Come Unstuck Why Gene Therapy Caused Leukemia in Some "Boy in the Bubble Syndrome" Patients Sugar Study is Sweetener for Stem Cell Science Insulin-Producing Cells Can Give Rise to Stem-Like Cells In-Vitro Standards in Stem Cell Research New Approach to Cancer: Find Most Tightly Controlled Genes Genetic Variation Increases HIV Risk in People of African Descent Genetic Profiling for Breast Cancer on the Horizon Doughnuts Deliver Drugs Stem Cell Therapies Offer Patients Hope and Reduced Health Care Costs Measuring the Footprint of Cells Adult Stem Cell Findings Offer New Hope for Parkinson's Cure Inhalable Form of Gene Therapy Takes Aim at Lung Cancer and Inflammatory Lung Disease Gene Therapy Involving Antibiotics May Help Patients With Usher Syndrome Discovery of New Family of Genetic Mutations Involved in Inflammatory Intestinal Disease Rapid Wound Healing With Dressing Made of Silica Gel Fibers Saving Teeth by Using Periodontal Ligament Regeneration Mapping of Prostate Cancer Genes Opens the Door to New Treatments Vancouver Researchers Pioneer Safe Pathway to Slow ALS Using Stem Cells Bone Repair Using Patient's Own Stem Cells Transfusable Blood From Stem Cells Pioneering Induction of Bone Formation Using Embryonic Stem Cells Muscle-Derived Stem Cells Prove Effective in Repairing Sphincter Damage to Restore Continence Discovery May Reverse/Prevent Cognitive Decline That Occurs During Ageing Dementia Genes Linked to Osteoporosis Identified Stem Cell Type Supposed to be Crucial for Angiogenesis and Cancer Growth Does Not Exist Cancer Cells Spread by Releasing 'Bubbles', According to New Study Tiny Magnets Used in Anti-Cancer Gene Therapy Stem Cell Hope for Fixing Injured Knees Clearer Day for Gene Therapy: New Vector Carries Big Genes Linked to Inherited Blindness Manchester University Announces Stem Cell Breakthrough New Venture for Therapy with Cord Blood and Adult Stem Cells World-First Stem Cell Screening Facility to Target Brain Tumors Way to Target Genes Devised for Gene Therapy Stem Cell Funding for Parkinsons Disease Stem Cell Breakthrough Offers Diabetes Hope Scottish Consortium to Start Work on 3D Cell-Based Assay Post Brain Injury: New Nerve Cells Originate From Neural Stem Cells Lifetime Probability of Needing a Stem Cell Transplant is Much Higher Than Previously Reported Deep-Brain Stimulation for Alzheimers Patients Considered Cheek Swab Kits Successful in Recruiting Bone Marrow Donors Breakthrough Technique Restores Light Sensitivity in Damaged Retinas Monash Pioneers Nano Scaffold to Rebuild Nerve Damage Stem Cell Therapy Trials to Mend Shattered Bones Coming Soon Patient Gets New Jaw Cultivated From Own Stem Cells Atlantic Veterinary College: New Technology Heals Horses Faster, Stronger Imitating Monkeys Jumping Genes Could Lead To New Treatments for HIV Human Clinical Trials of Stem Cell Treatment That Could Replace Bone Marrow Transplants New Acute Lung Injury Study Finds Benefits of Autologous Cell Transplantation Therapy Helmet Helps Alzheimers Patients Polymer Gel Prevents Skin Grafts from Shrinking 'Telepathic' Genes Recognize Similarities in Each Other Discovery: Mutations of a Part of the X Chromosome Lead to Mental Retardation Treatment for Brittle Bones Before Birth Twins Confirm the Beginning of Childhood Leukemia Macular Degeneration Stem Cell Therapy Repair May Be On the Horizon Lab-Grown Brain Cells Could Treat Parkinsons Disease Gene Suppression Enhances Wound Healing Research on Treatments to Improve Muscle Loss Due to Disease and Aging Gene Therapy Experiences Another Setback in UK Growing Artificial Skin From Hair Roots Gene Therapy May Protect Against Alcoholism Regenerative Medicine Researchers Use Fat to Repair Breast Defects Following a Lumpectomy Regenerative Medicine Aid: New Ultrasound Device Determines Best Time for Skin Transplant Muscular Dystrophy May Be Helped Through Regenerative Medicine Therapy Face Transplant Patient Doing Well Today Regenerative Stem Cell Therapy Helps Memory New Source for Dopamine Cells May Help Those With Parkinsons Disease Important Step in Development of Artificial Nerves via Regenerative Medicine Endometrial Regenerative Cells: New Adult Stem Cells for Regenerative Medicine Adult Stem Cell Product to be Tested in Regenerative Medicine Stroke Trial New Regenerative Medicine Stem Cell Treatment Reduces Amputations in Critical Limb Ischemia Using Stem Cells to Treat Damaged Eyes and a Rare Skin Disorder Regenerative Medicine Efforts Honored: 2007 Prix DExcellence Award Inauguration of the Clinical Translation Unit Treatment of Vascular Risk Factors Slows Advance of Alzheimers Skin-Derived Stem Cells Repair Spinal Cord Injuries in Rats Gene Therapy in Rats Reduces Rheumatoid Arthritis Swelling Growing Heart Valves Novalung: Bridge to Lung Transplant Stem Cell Subtype Discovery Could Lead to Improved Bone Marrow Transplants Bone-Strengthening Drugs May Benefit Young Stem Cell Transplant Recipients "Off-the-Shelf" Skin Grafts Stem Cells Grow Kidneys in Mice New Source for Stem Cell Line One For the Road: An Alcoholics Last Shot? Discovery Could Lead to Regenerative Therapies for Muscular Disease Gene Therapy Works to Eradicate Bowel Cancer Cells Gene Therapy Trial for Treatment of Lebers Congenital Amaurosis (LCA) Bone Marrow May Help With Future Male Fertility Treatments Stem Cell Treatment for Runners Stem Cell Treatment for Recently Diagnosed Type 1 Diabetics Liver Cells Grown From Cord Blood© McGowan Institute for Regenerative Medicine | Contact Us